

You can:
| Name | Lysophosphatidic acid receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | LPAR2 |
| Synonym | endothelial differentiation gene 4, lysophosphatidic acid G-protein-coupled receptor 4 LPA receptor 2 LPA-2 Edg4 Lysophosphatidic acid receptor Edg-4 [ Show all ] |
| Disease | N/A |
| Length | 351 |
| Amino acid sequence | MVIMGQCYYNETIGFFYNNSGKELSSHWRPKDVVVVALGLTVSVLVLLTNLLVIAAIASNRRFHQPIYYLLGNLAAADLFAGVAYLFLMFHTGPRTARLSLEGWFLRQGLLDTSLTASVATLLAIAVERHRSVMAVQLHSRLPRGRVVMLIVGVWVAALGLGLLPAHSWHCLCALDRCSRMAPLLSRSYLAVWALSSLLVFLLMVAVYTRIFFYVRRRVQRMAEHVSCHPRYRETTLSLVKTVVIILGAFVVCWTPGQVVLLLDGLGCESCNVLAVEKYFLLLAEANSLVNAAVYSCRDAEMRRTFRRLLCCACLRQSTRESVHYTSSAQGGASTRIMLPENGHPLMDSTL |
| UniProt | Q9HBW0 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9HBW0 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9HBW0. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3724 |
| IUPHAR | 273 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3426 | 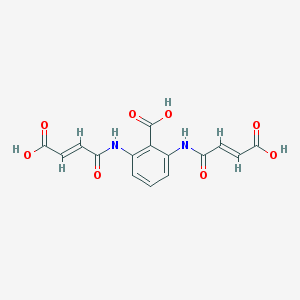 CHEMBL485119 CHEMBL485119 | C15H12N2O8 | 348.267 | 8 / 5 | -0.1 | Yes |
| 8958 | 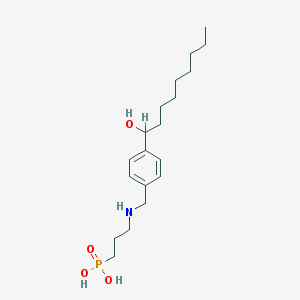 CHEMBL117007 CHEMBL117007 | C19H34NO4P | 371.458 | 5 / 4 | 0.6 | Yes |
| 16494 | 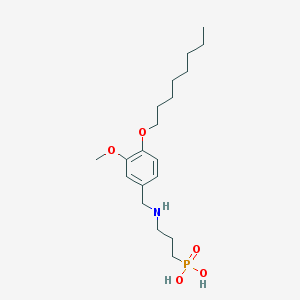 CHEMBL118508 CHEMBL118508 | C19H34NO5P | 387.457 | 6 / 3 | 0.8 | Yes |
| 17534 | 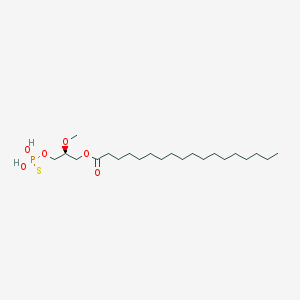 CHEMBL357053 CHEMBL357053 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17536 | 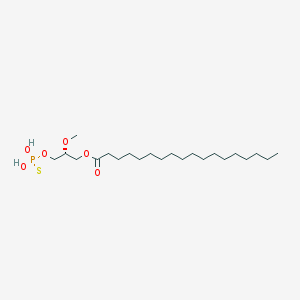 CHEMBL153043 CHEMBL153043 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17543 | 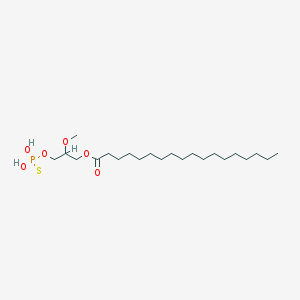 CHEMBL2017139 CHEMBL2017139 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 20483 | 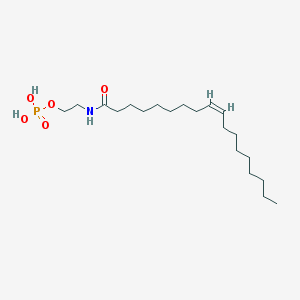 NAEPA NAEPA | C20H40NO5P | 405.516 | 5 / 3 | 5.2 | No |
| 21364 | 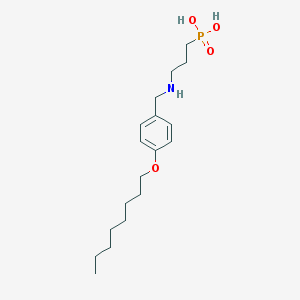 CHEMBL332373 CHEMBL332373 | C18H32NO4P | 357.431 | 5 / 3 | 0.9 | Yes |
| 442567 | 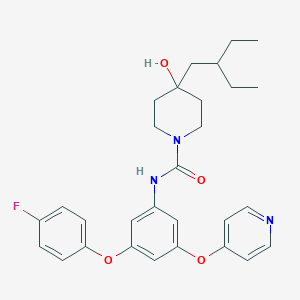 CHEMBL3401388 CHEMBL3401388 | C29H34FN3O4 | 507.606 | 6 / 2 | 5.7 | No |
| 25664 | 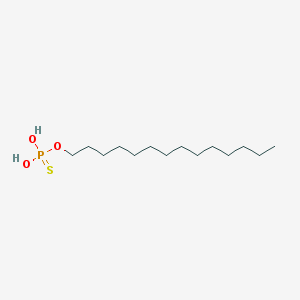 CHEMBL189296 CHEMBL189296 | C14H31O3PS | 310.433 | 4 / 2 | 6.8 | No |
| 27911 | 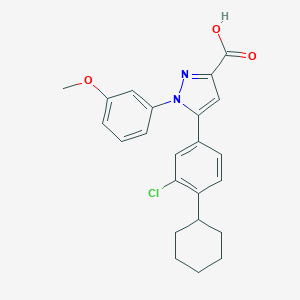 TC LPA5 4 TC LPA5 4 | C23H23ClN2O3 | 410.898 | 4 / 1 | 6.5 | No |
| 29831 | 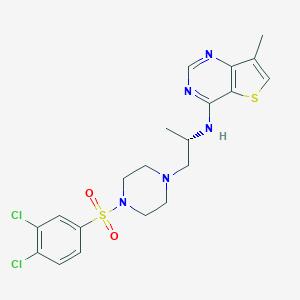 LPA2 antagonist 1 LPA2 antagonist 1 | C20H23Cl2N5O2S2 | 500.457 | 8 / 1 | 4.3 | No |
| 467010 | 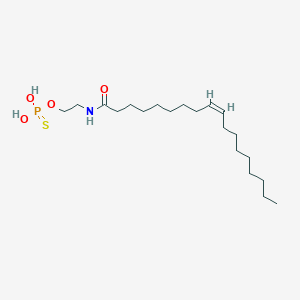 CHEMBL3621959 CHEMBL3621959 | C20H40NO4PS | 421.577 | 5 / 3 | 7.0 | No |
| 33972 | 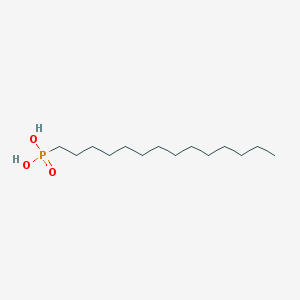 Tetradecylphosphonic acid Tetradecylphosphonic acid | C14H31O3P | 278.373 | 3 / 2 | 5.1 | No |
| 34949 | 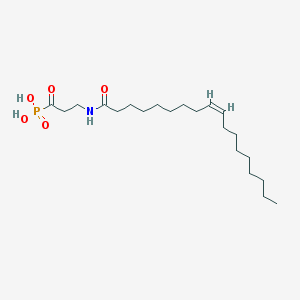 CHEMBL117529 CHEMBL117529 | C21H40NO5P | 417.527 | 5 / 3 | 4.9 | Yes |
| 35908 | 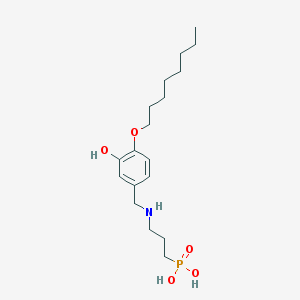 CHEMBL119239 CHEMBL119239 | C18H32NO5P | 373.43 | 6 / 4 | 0.6 | Yes |
| 37810 | 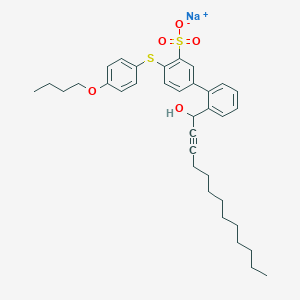 CHEMBL239659 CHEMBL239659 | C35H43NaO5S2 | 630.834 | 6 / 1 | N/A | No |
| 443332 | 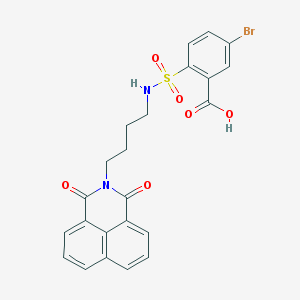 CHEMBL3322508 CHEMBL3322508 | C23H19BrN2O6S | 531.377 | 7 / 2 | 3.7 | No |
| 443441 | 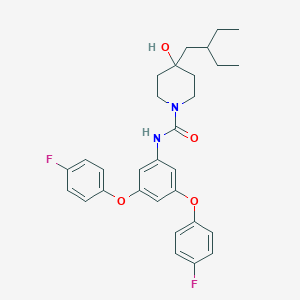 CHEMBL3401379 CHEMBL3401379 | C30H34F2N2O4 | 524.609 | 6 / 2 | 6.8 | No |
| 47585 | 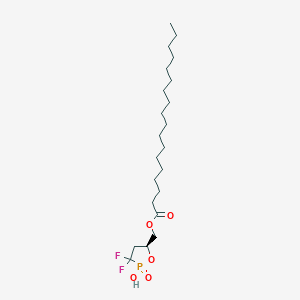 CHEMBL215722 CHEMBL215722 | C22H41F2O5P | 454.536 | 7 / 1 | 8.2 | No |
| 51246 | 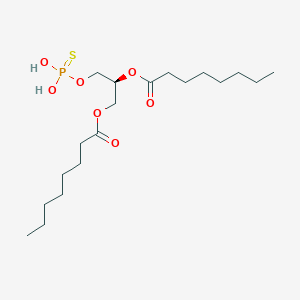 CHEMBL202242 CHEMBL202242 | C19H37O7PS | 440.532 | 8 / 2 | 6.0 | No |
| 51247 | 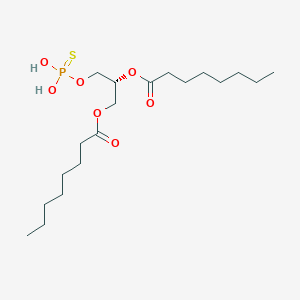 CHEMBL202361 CHEMBL202361 | C19H37O7PS | 440.532 | 8 / 2 | 6.0 | No |
| 537351 | 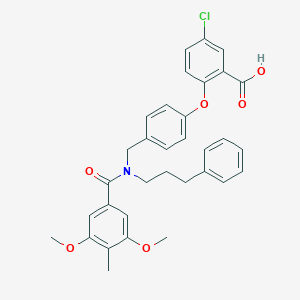 CHEMBL3975893 CHEMBL3975893 | C33H32ClNO6 | 574.07 | 6 / 1 | 7.2 | No |
| 65691 | 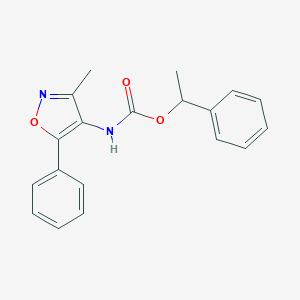 CHEMBL404575 CHEMBL404575 | C19H18N2O3 | 322.364 | 4 / 1 | 3.9 | Yes |
| 66740 | 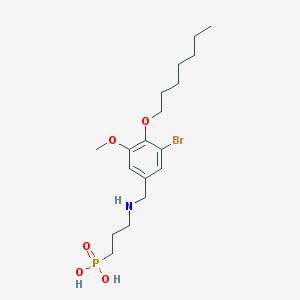 CHEMBL118783 CHEMBL118783 | C18H31BrNO5P | 452.326 | 6 / 3 | 1.1 | Yes |
| 67557 | 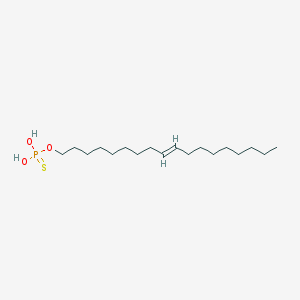 CHEMBL187459 CHEMBL187459 | C18H37O3PS | 364.525 | 4 / 2 | 8.1 | No |
| 69973 | 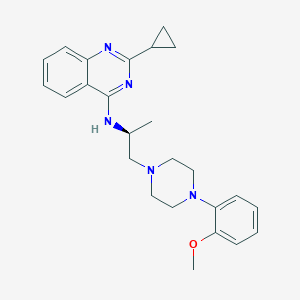 CHEMBL270878 CHEMBL270878 | C25H31N5O | 417.557 | 6 / 1 | 4.5 | Yes |
| 71481 | 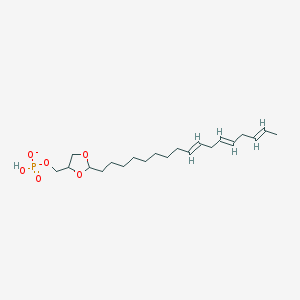 CHEMBL383362 CHEMBL383362 | C21H36O6P- | 415.487 | 6 / 1 | 4.7 | Yes |
| 75847 | 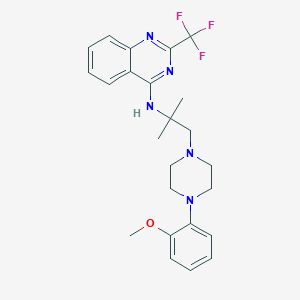 CHEMBL257899 CHEMBL257899 | C24H28F3N5O | 459.517 | 9 / 1 | 5.0 | Yes |
| 85970 | 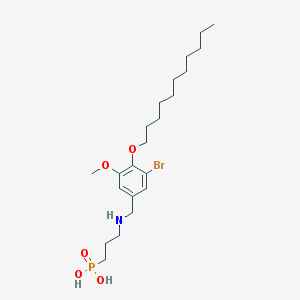 CHEMBL118815 CHEMBL118815 | C22H39BrNO5P | 508.434 | 6 / 3 | 3.2 | No |
| 86133 | 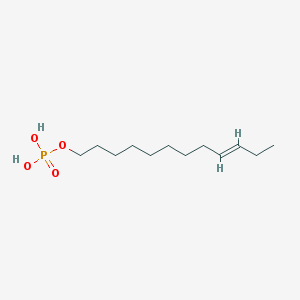 CHEMBL190349 CHEMBL190349 | C12H25O4P | 264.302 | 4 / 2 | 3.1 | Yes |
| 86861 | 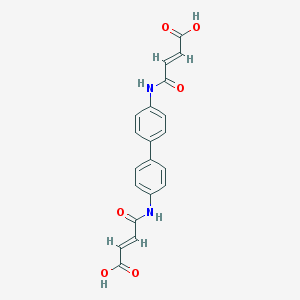 36840-10-5 36840-10-5 | C20H16N2O6 | 380.356 | 6 / 4 | 1.4 | Yes |
| 88209 | 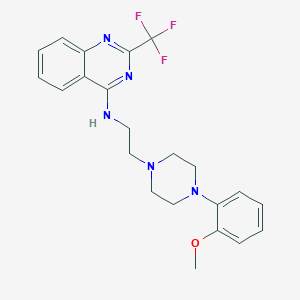 CHEMBL272352 CHEMBL272352 | C22H24F3N5O | 431.463 | 9 / 1 | 4.4 | Yes |
| 91232 | 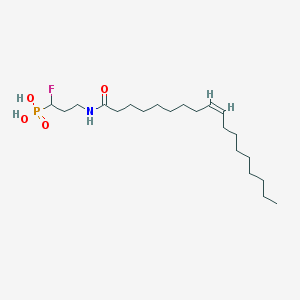 CHEMBL117663 CHEMBL117663 | C21H41FNO4P | 421.534 | 5 / 3 | 6.1 | No |
| 92112 | 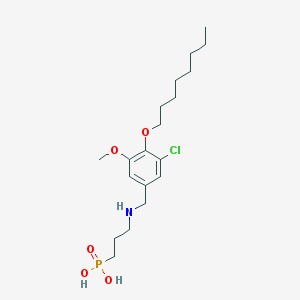 CHEMBL117715 CHEMBL117715 | C19H33ClNO5P | 421.899 | 6 / 3 | 1.5 | Yes |
| 93260 | 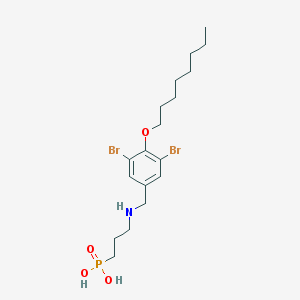 CHEMBL332667 CHEMBL332667 | C18H30Br2NO4P | 515.223 | 5 / 3 | 2.3 | No |
| 445646 | 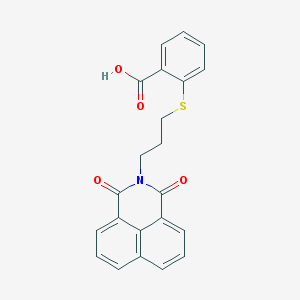 325850-81-5 325850-81-5 | C22H17NO4S | 391.441 | 5 / 1 | 4.2 | Yes |
| 101764 | 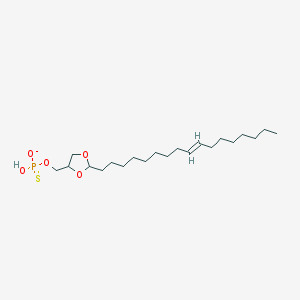 CHEMBL199472 CHEMBL199472 | C21H40O5PS- | 435.58 | 6 / 1 | 7.7 | No |
| 101768 | 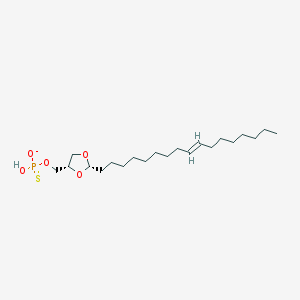 CHEMBL425218 CHEMBL425218 | C21H40O5PS- | 435.58 | 6 / 1 | 7.7 | No |
| 101769 | 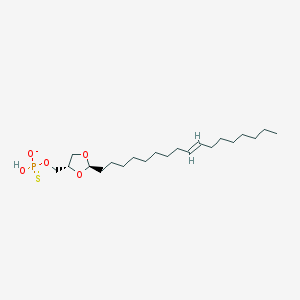 CHEMBL200003 CHEMBL200003 | C21H40O5PS- | 435.58 | 6 / 1 | 7.7 | No |
| 101772 | 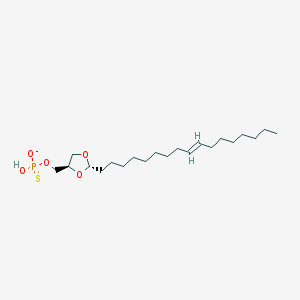 CHEMBL427014 CHEMBL427014 | C21H40O5PS- | 435.58 | 6 / 1 | 7.7 | No |
| 101777 | 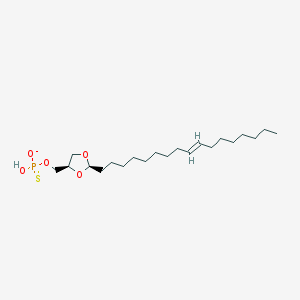 CHEMBL199400 CHEMBL199400 | C21H40O5PS- | 435.58 | 6 / 1 | 7.7 | No |
| 102473 | 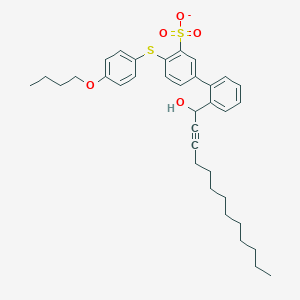 CHEMBL239659 CHEMBL239659 | C35H43O5S2- | 607.844 | 6 / 1 | 10.2 | No |
| 445760 | 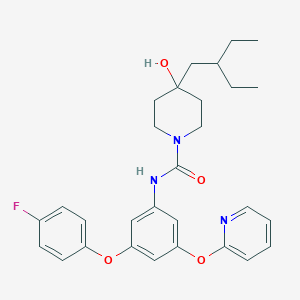 CHEMBL3401390 CHEMBL3401390 | C29H34FN3O4 | 507.606 | 6 / 2 | 6.0 | No |
| 445833 | 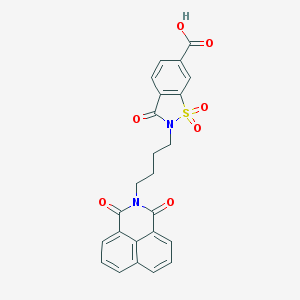 CHEMBL3322509 CHEMBL3322509 | C24H18N2O7S | 478.475 | 7 / 1 | 2.8 | Yes |
| 109233 | 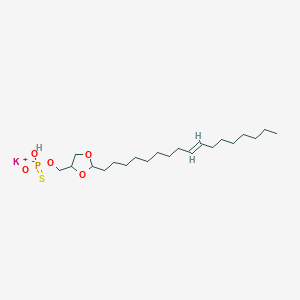 CHEMBL199472 CHEMBL199472 | C21H40KO5PS | 474.678 | 6 / 1 | N/A | N/A |
| 109236 | 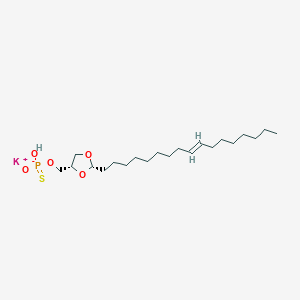 CHEMBL425218 CHEMBL425218 | C21H40KO5PS | 474.678 | 6 / 1 | N/A | N/A |
| 109240 | 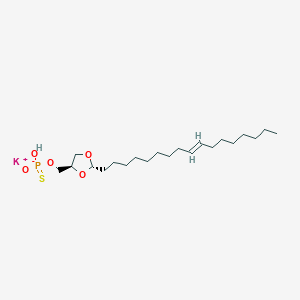 CHEMBL427014 CHEMBL427014 | C21H40KO5PS | 474.678 | 6 / 1 | N/A | N/A |
| 109244 | 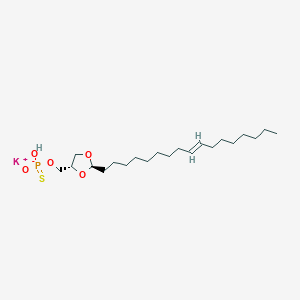 CHEMBL200003 CHEMBL200003 | C21H40KO5PS | 474.678 | 6 / 1 | N/A | N/A |
| 109247 | 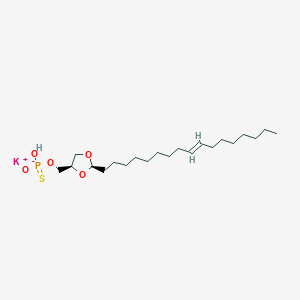 CHEMBL199400 CHEMBL199400 | C21H40KO5PS | 474.678 | 6 / 1 | N/A | N/A |
| 121712 | 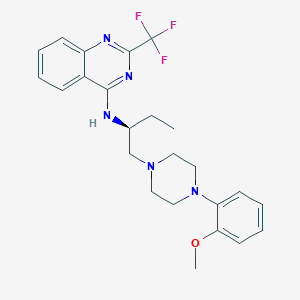 CHEMBL406676 CHEMBL406676 | C24H28F3N5O | 459.517 | 9 / 1 | 5.3 | No |
| 122120 | 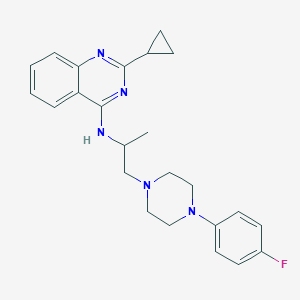 CHEMBL402218 CHEMBL402218 | C24H28FN5 | 405.521 | 6 / 1 | 4.6 | Yes |
| 126104 | 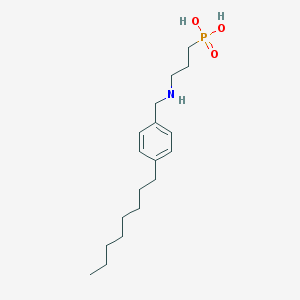 CHEMBL119256 CHEMBL119256 | C18H32NO3P | 341.432 | 4 / 3 | 1.6 | Yes |
| 127640 | 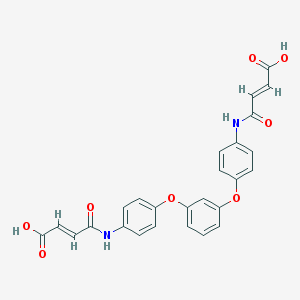 CHEMBL482498 CHEMBL482498 | C26H20N2O8 | 488.452 | 8 / 4 | 2.9 | Yes |
| 129052 | 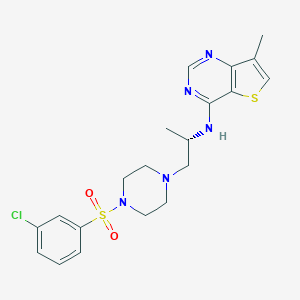 CHEMBL270865 CHEMBL270865 | C20H24ClN5O2S2 | 466.015 | 8 / 1 | 3.7 | Yes |
| 133531 | 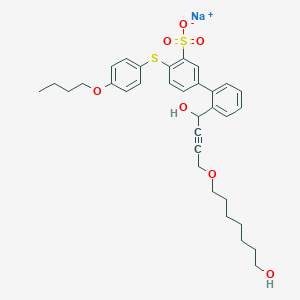 CHEMBL397081 CHEMBL397081 | C33H39NaO7S2 | 634.778 | 8 / 2 | N/A | No |
| 140724 | 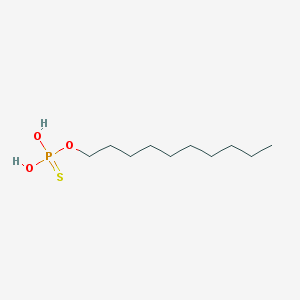 Thiophosphoric acid decyl ester Thiophosphoric acid decyl ester | C10H23O3PS | 254.325 | 4 / 2 | 4.7 | Yes |
| 141052 | 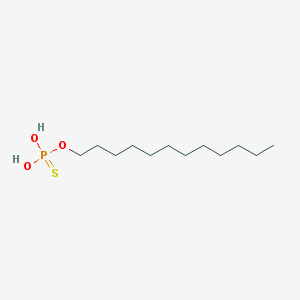 dodecyl-thiophosphate dodecyl-thiophosphate | C12H27O3PS | 282.379 | 4 / 2 | 5.8 | No |
| 144860 | 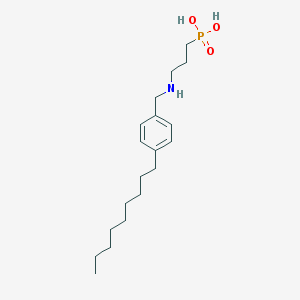 CHEMBL118860 CHEMBL118860 | C19H34NO3P | 355.459 | 4 / 3 | 2.1 | Yes |
| 554000 | 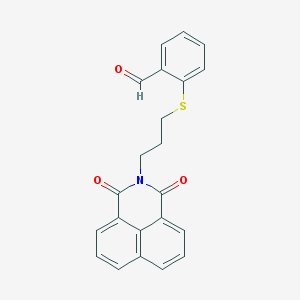 D0G8FF D0G8FF | C22H17NO3S | 375.442 | 4 / 0 | 4.2 | Yes |
| 151960 | 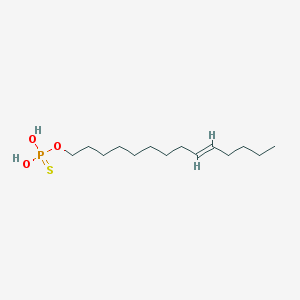 CHEMBL187402 CHEMBL187402 | C14H29O3PS | 308.417 | 4 / 2 | 5.9 | No |
| 154929 | 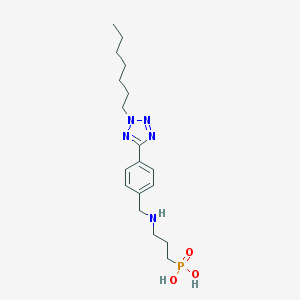 CHEMBL119382 CHEMBL119382 | C18H30N5O3P | 395.444 | 7 / 3 | 0.2 | Yes |
| 162855 | 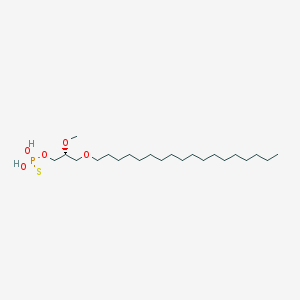 CHEMBL2335048 CHEMBL2335048 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162865 | 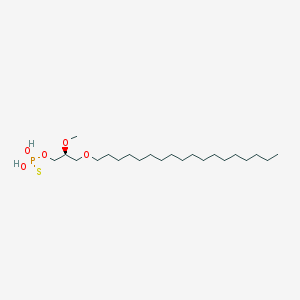 CHEMBL2335047 CHEMBL2335047 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162867 | 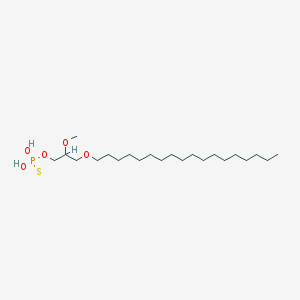 CHEMBL2335052 CHEMBL2335052 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 163619 | 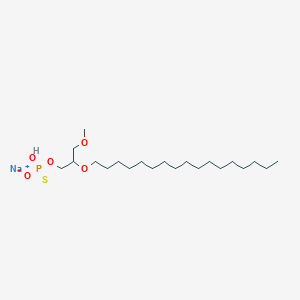 CHEMBL2335050 CHEMBL2335050 | C21H44NaO5PS | 462.602 | 6 / 1 | N/A | N/A |
| 164314 | 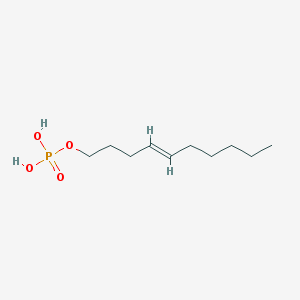 CHEMBL188081 CHEMBL188081 | C10H21O4P | 236.248 | 4 / 2 | 2.2 | Yes |
| 176842 | 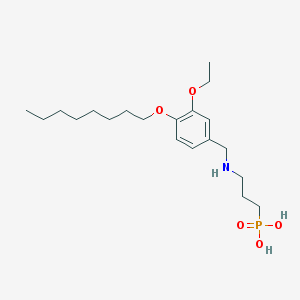 CHEMBL324820 CHEMBL324820 | C20H36NO5P | 401.484 | 6 / 3 | 1.3 | Yes |
| 178100 | 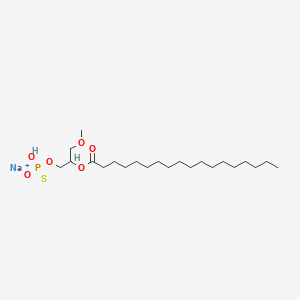 CHEMBL2335051 CHEMBL2335051 | C22H44NaO6PS | 490.612 | 7 / 1 | N/A | N/A |
| 554223 | 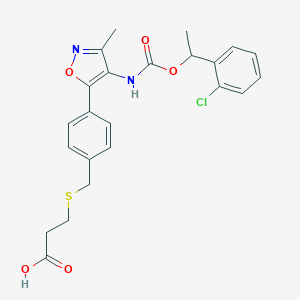 Ki16425 Ki16425 | C23H23ClN2O5S | 474.956 | 7 / 2 | 4.5 | Yes |
| 486637 | 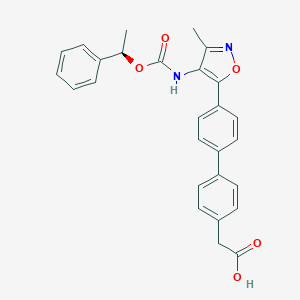 AM095 free acid AM095 free acid | C27H24N2O5 | 456.498 | 6 / 2 | 5.0 | Yes |
| 197612 | 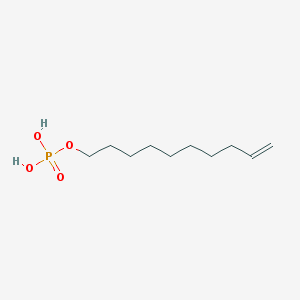 Phosphoric acid monodec-9-enyl ester Phosphoric acid monodec-9-enyl ester | C10H21O4P | 236.248 | 4 / 2 | 2.4 | Yes |
| 198414 | 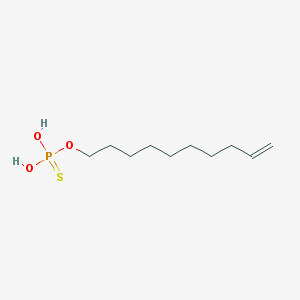 Thiophosphoric acid dec-9-enyl ester Thiophosphoric acid dec-9-enyl ester | C10H21O3PS | 252.309 | 4 / 2 | 4.2 | Yes |
| 199821 | 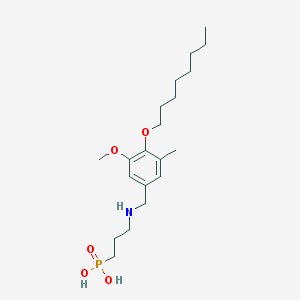 CHEMBL432813 CHEMBL432813 | C20H36NO5P | 401.484 | 6 / 3 | 1.3 | Yes |
| 209042 | 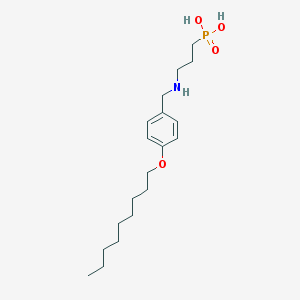 CHEMBL333335 CHEMBL333335 | C19H34NO4P | 371.458 | 5 / 3 | 1.5 | Yes |
| 217785 | 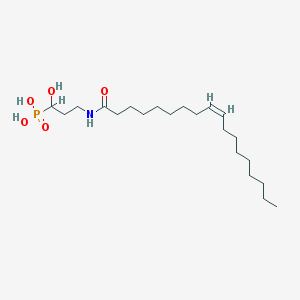 SCHEMBL3448131 SCHEMBL3448131 | C21H42NO5P | 419.543 | 5 / 4 | 5.0 | Yes |
| 450306 | 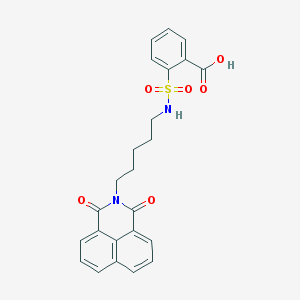 CHEMBL3322503 CHEMBL3322503 | C24H22N2O6S | 466.508 | 7 / 2 | 3.4 | Yes |
| 230380 | 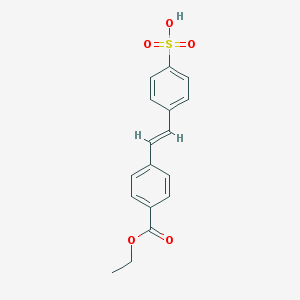 CHEMBL483303 CHEMBL483303 | C17H16O5S | 332.37 | 5 / 1 | 3.2 | Yes |
| 230720 | 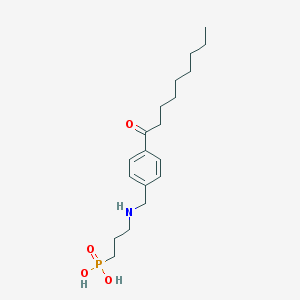 CHEMBL115970 CHEMBL115970 | C19H32NO4P | 369.442 | 5 / 3 | 0.8 | Yes |
| 236633 | 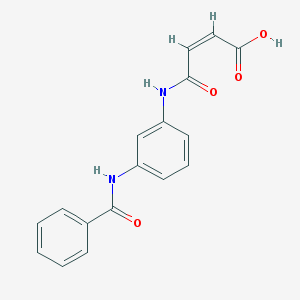 CHEMBL261872 CHEMBL261872 | C17H14N2O4 | 310.309 | 4 / 3 | 1.6 | Yes |
| 241086 | 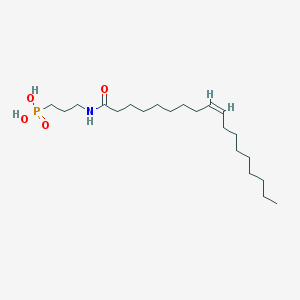 CHEMBL331661 CHEMBL331661 | C21H42NO4P | 403.544 | 4 / 3 | 5.5 | No |
| 451312 | 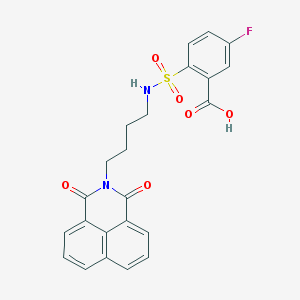 CHEMBL3322513 CHEMBL3322513 | C23H19FN2O6S | 470.471 | 8 / 2 | 3.2 | Yes |
| 451414 | 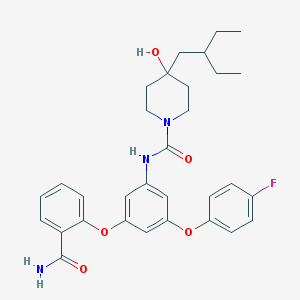 CHEMBL3401385 CHEMBL3401385 | C31H36FN3O5 | 549.643 | 6 / 3 | 5.6 | No |
| 246831 | 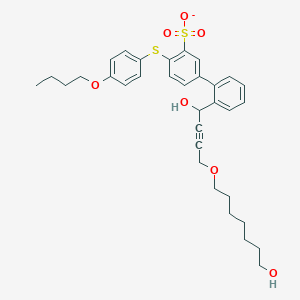 CHEMBL397081 CHEMBL397081 | C33H39O7S2- | 611.788 | 8 / 2 | 6.0 | No |
| 246947 | 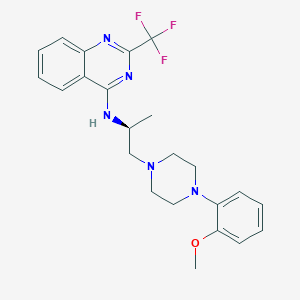 CHEMBL270877 CHEMBL270877 | C23H26F3N5O | 445.49 | 9 / 1 | 4.8 | Yes |
| 246948 | 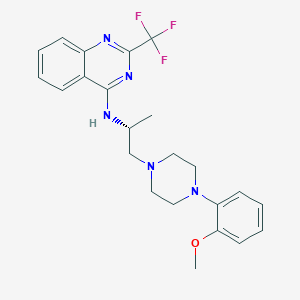 CHEMBL256478 CHEMBL256478 | C23H26F3N5O | 445.49 | 9 / 1 | 4.8 | Yes |
| 451615 | 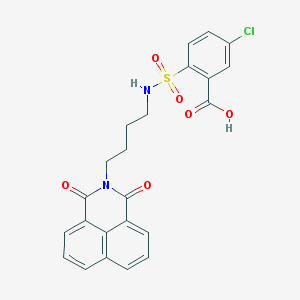 CHEMBL3322514 CHEMBL3322514 | C23H19ClN2O6S | 486.923 | 7 / 2 | 3.7 | Yes |
| 256134 | 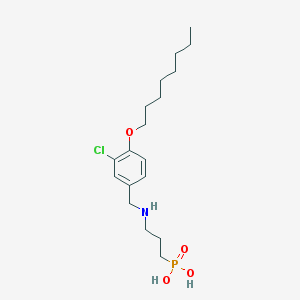 CHEMBL119413 CHEMBL119413 | C18H31ClNO4P | 391.873 | 5 / 3 | 1.6 | Yes |
| 451813 | 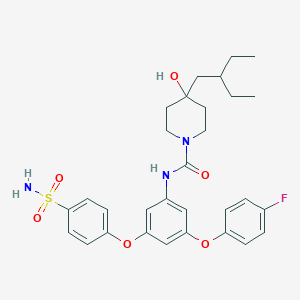 CHEMBL3401382 CHEMBL3401382 | C30H36FN3O6S | 585.691 | 8 / 3 | 5.3 | No |
| 262446 | 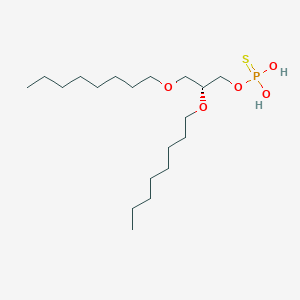 CHEMBL201482 CHEMBL201482 | C19H41O5PS | 412.566 | 6 / 2 | 6.8 | No |
| 542968 | 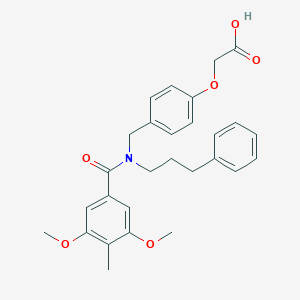 CHEMBL3956307 CHEMBL3956307 | C28H31NO6 | 477.557 | 6 / 1 | 5.1 | No |
| 452175 | 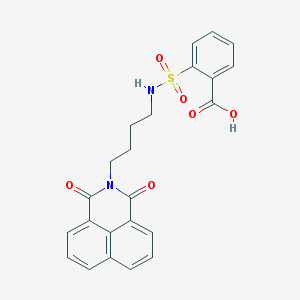 DBIBB DBIBB | C23H20N2O6S | 452.481 | 7 / 2 | 3.1 | Yes |
| 264962 | 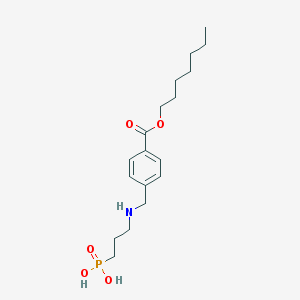 CHEMBL117403 CHEMBL117403 | C18H30NO5P | 371.414 | 6 / 3 | 0.3 | Yes |
| 269089 | 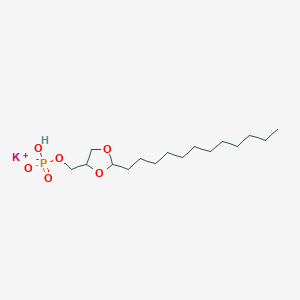 CHEMBL381470 CHEMBL381470 | C16H32KO6P | 390.498 | 6 / 1 | N/A | N/A |
| 277526 | 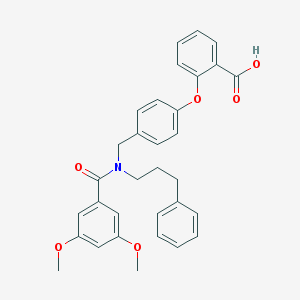 CHEMBL272087 CHEMBL272087 | C32H31NO6 | 525.601 | 6 / 1 | 6.2 | No |
| 278363 | 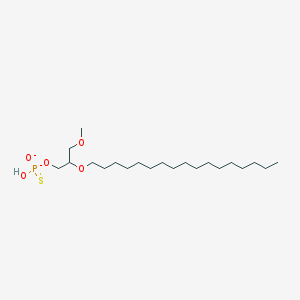 CHEMBL2335050 CHEMBL2335050 | C21H44O5PS- | 439.612 | 6 / 1 | 8.2 | No |
| 278838 | 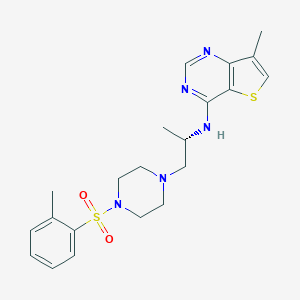 CHEMBL429681 CHEMBL429681 | C21H27N5O2S2 | 445.6 | 8 / 1 | 3.4 | Yes |
| 290479 | 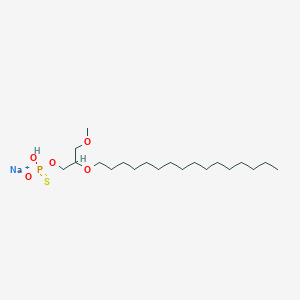 CHEMBL2335049 CHEMBL2335049 | C20H42NaO5PS | 448.575 | 6 / 1 | N/A | N/A |
| 293491 | 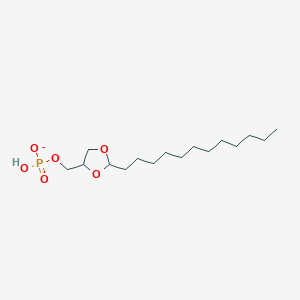 CHEMBL381470 CHEMBL381470 | C16H32O6P- | 351.4 | 6 / 1 | 4.2 | Yes |
| 305407 | 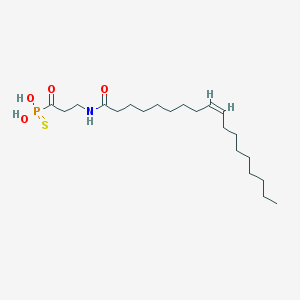 CHEMBL117798 CHEMBL117798 | C21H40NO4PS | 433.588 | 5 / 3 | 6.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218