

You can:
| Name | Leukotriene B4 receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | LTB4R2 |
| Synonym | NOP9 12-HHT receptor Seven transmembrane receptor BLTR2 LTB4-R2 LTB4-R 2 [ Show all ] |
| Disease | Inflammatory bowel disease |
| Length | 358 |
| Amino acid sequence | MSVCYRPPGNETLLSWKTSRATGTAFLLLAALLGLPGNGFVVWSLAGWRPARGRPLAATLVLHLALADGAVLLLTPLFVAFLTRQAWPLGQAGCKAVYYVCALSMYASVLLTGLLSLQRCLAVTRPFLAPRLRSPALARRLLLAVWLAALLLAVPAAVYRHLWRDRVCQLCHPSPVHAAAHLSLETLTAFVLPFGLMLGCYSVTLARLRGARWGSGRHGARVGRLVSAIVLAFGLLWAPYHAVNLLQAVAALAPPEGALAKLGGAGQAARAGTTALAFFSSSVNPVLYVFTAGDLLPRAGPRFLTRLFEGSGEARGGGRSREGTMELRTTPQLKVVGQGRGNGDPGGGMEKDGPEWDL |
| UniProt | Q9NPC1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9NPC1 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9NPC1. |
| BioLiP | N/A |
| Therapeutic Target Database | T30563 |
| ChEMBL | CHEMBL3191 |
| IUPHAR | 268 |
| DrugBank | BE0003489 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 459278 | 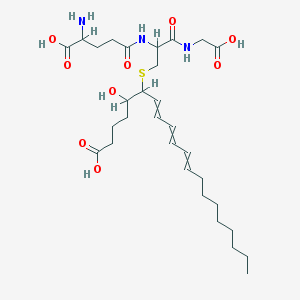 Leukotriene C3 Leukotriene C3 | C30H49N3O9S | 627.794 | 11 / 7 | 1.3 | No |
| 24689 | 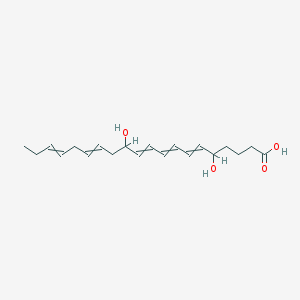 5,12-dihydroxyicosa-6,8,10,14,17-pentaenoic acid 5,12-dihydroxyicosa-6,8,10,14,17-pentaenoic acid | C20H30O4 | 334.456 | 4 / 3 | 3.4 | Yes |
| 555642 | 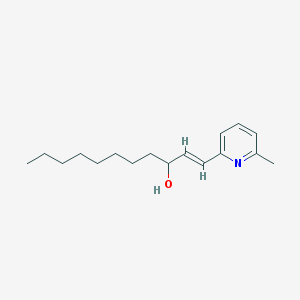 BDBM81523 BDBM81523 | C17H27NO | 261.409 | 2 / 1 | 5.0 | Yes |
| 555675 | 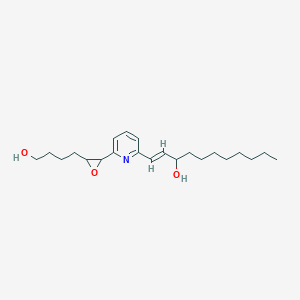 BDBM81520 BDBM81520 | C22H35NO3 | 361.526 | 4 / 2 | 4.6 | Yes |
| 555740 | 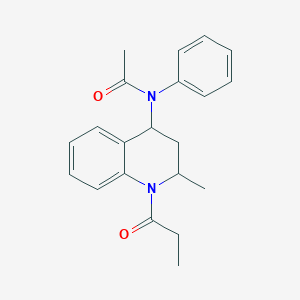 Probes1_000120 Probes1_000120 | C21H24N2O2 | 336.435 | 2 / 0 | 3.2 | Yes |
| 555789 | 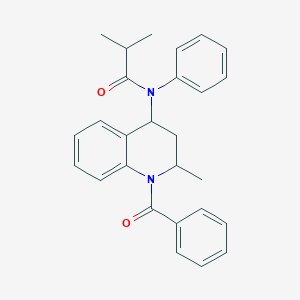 2-methyl-N-[2-methyl-1-(phenylcarbonyl)-1,2,3,4-tetrahydroquinolin-4-yl]-N-phenylpropanamide 2-methyl-N-[2-methyl-1-(phenylcarbonyl)-1,2,3,4-tetrahydroquinolin-4-yl]-N-phenylpropanamide | C27H28N2O2 | 412.533 | 2 / 0 | 5.4 | No |
| 97414 | 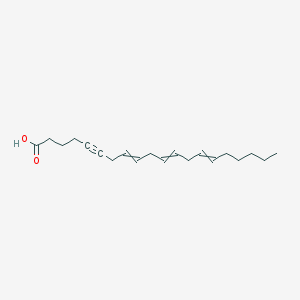 8,11,14-eicosatrien-5-ynoic acid 8,11,14-eicosatrien-5-ynoic acid | C20H30O2 | 302.458 | 2 / 1 | 6.0 | No |
| 105854 | 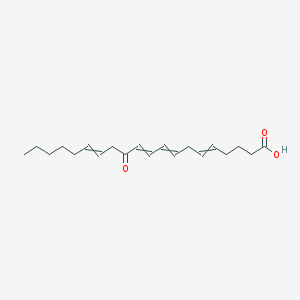 5,8,10,14-Eicosatetraenoicacid, 12-oxo-, (5Z,8Z,10E,14Z)- 5,8,10,14-Eicosatetraenoicacid, 12-oxo-, (5Z,8Z,10E,14Z)- | C20H30O3 | 318.457 | 3 / 1 | 5.1 | No |
| 107099 | 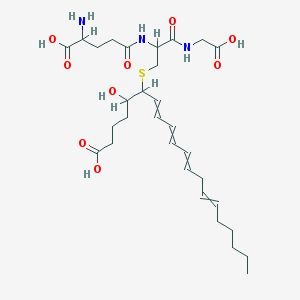 LTC4 (Leukotriene C4) LTC4 (Leukotriene C4) | C30H47N3O9S | 625.778 | 11 / 7 | 0.7 | No |
| 111129 | 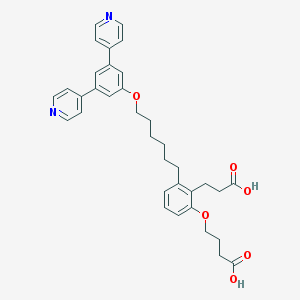 CHEMBL1099326 CHEMBL1099326 | C35H38N2O6 | 582.697 | 8 / 2 | 6.3 | No |
| 553858 | 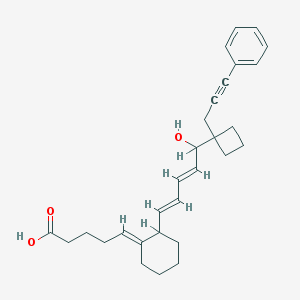 ZK158252 ZK158252 | C29H36O3 | 432.604 | 3 / 2 | 6.4 | No |
| 555943 | 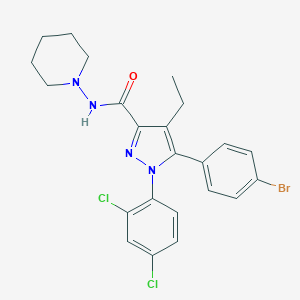 Surinabant Surinabant | C23H23BrCl2N4O | 522.268 | 3 / 1 | 7.0 | No |
| 139430 | 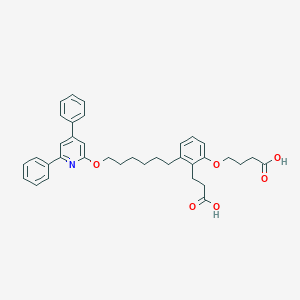 CHEMBL1099330 CHEMBL1099330 | C36H39NO6 | 581.709 | 7 / 2 | 7.8 | No |
| 553989 | 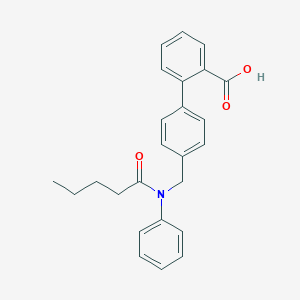 CAY10583 CAY10583 | C25H25NO3 | 387.479 | 3 / 1 | 5.3 | No |
| 143814 | 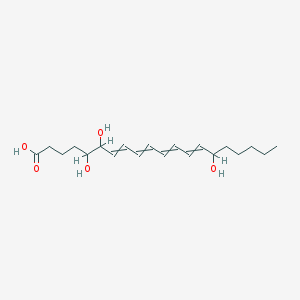 5,6,15-trihydroxyicosa-7,9,11,13-tetraenoic acid 5,6,15-trihydroxyicosa-7,9,11,13-tetraenoic acid | C20H32O5 | 352.471 | 5 / 4 | 3.1 | Yes |
| 460617 | 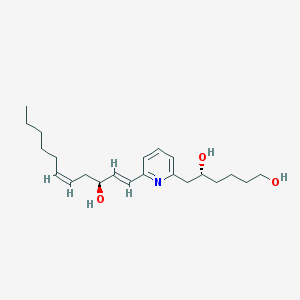 MLS000028864 MLS000028864 | C22H35NO3 | 361.526 | 4 / 3 | 4.1 | Yes |
| 154967 | 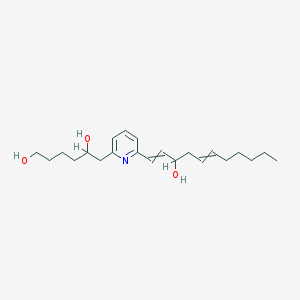 U-75302 U-75302 | C22H35NO3 | 361.526 | 4 / 3 | 4.1 | Yes |
| 556067 | 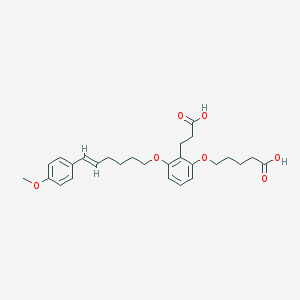 ONO-4057 ONO-4057 | C27H34O7 | 470.562 | 7 / 2 | 5.1 | No |
| 460651 | 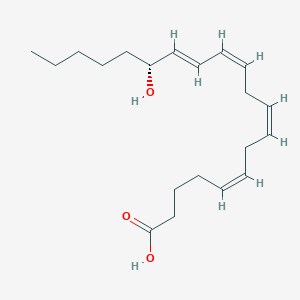 15R-HETE 15R-HETE | C20H32O3 | 320.473 | 3 / 2 | 5.1 | No |
| 158686 | 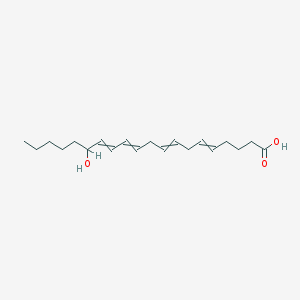 15(S)-HETE-d8 15(S)-HETE-d8 | C20H32O3 | 320.473 | 3 / 2 | 5.1 | No |
| 554077 | 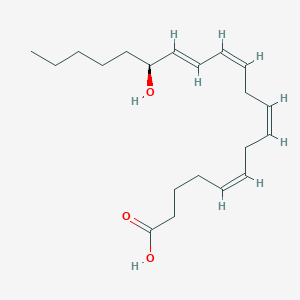 Icomucret Icomucret | C20H32O3 | 320.473 | 3 / 2 | 5.1 | No |
| 168508 | 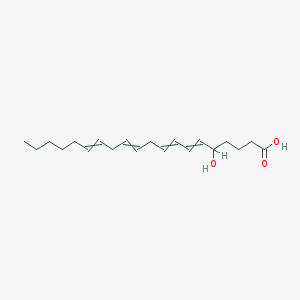 5-Hete 5-Hete | C20H32O3 | 320.473 | 3 / 2 | 5.2 | No |
| 460720 | 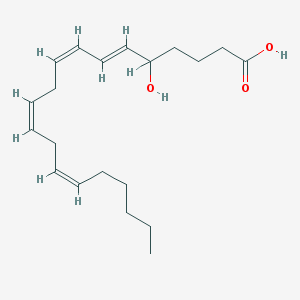 5-Hete 5-Hete | C20H32O3 | 320.473 | 3 / 2 | 5.2 | No |
| 460792 | 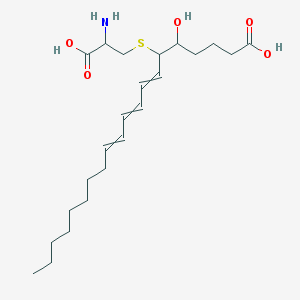 CTK9A5157 CTK9A5157 | C23H39NO5S | 441.627 | 7 / 4 | 2.8 | Yes |
| 554179 | 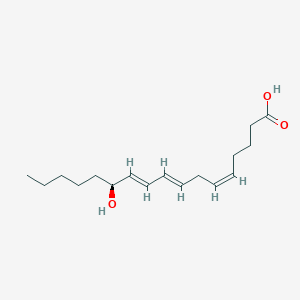 12S-HHTrE 12S-HHTrE | C17H28O3 | 280.408 | 3 / 2 | 4.1 | Yes |
| 203201 | 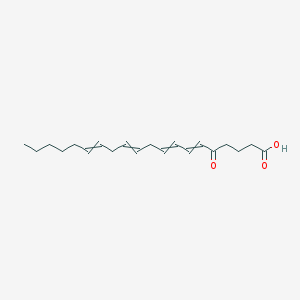 6,8,11,14-Eicosatetraenoicacid, 5-oxo- 6,8,11,14-Eicosatetraenoicacid, 5-oxo- | C20H30O3 | 318.457 | 3 / 1 | 5.1 | No |
| 556295 | 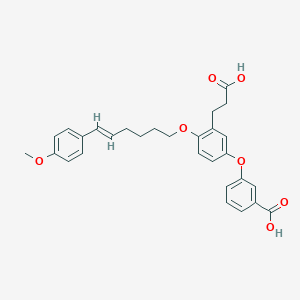 BDBM81522 BDBM81522 | C29H30O7 | 490.552 | 7 / 2 | 6.0 | No |
| 226540 | 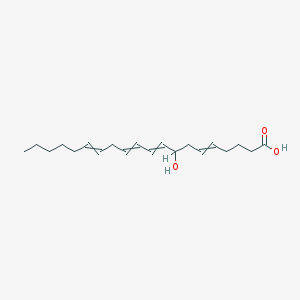 8-hydroxyicosa-5,9,11,14-tetraenoic acid 8-hydroxyicosa-5,9,11,14-tetraenoic acid | C20H32O3 | 320.473 | 3 / 2 | 5.2 | No |
| 227070 | 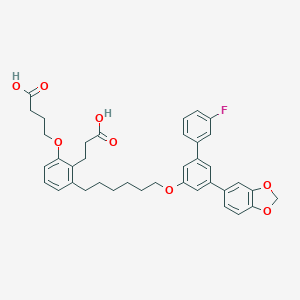 CHEMBL1094348 CHEMBL1094348 | C38H39FO8 | 642.72 | 9 / 2 | 8.4 | No |
| 227534 | 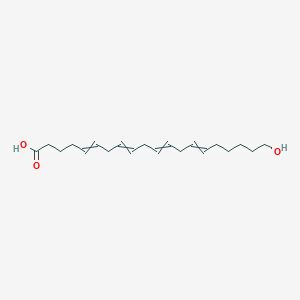 5,8,11,14-Eicosatetraenoicacid, 20-hydroxy-, (5Z,8Z,11Z,14Z)- 5,8,11,14-Eicosatetraenoicacid, 20-hydroxy-, (5Z,8Z,11Z,14Z)- | C20H32O3 | 320.473 | 3 / 2 | 4.7 | Yes |
| 554449 | 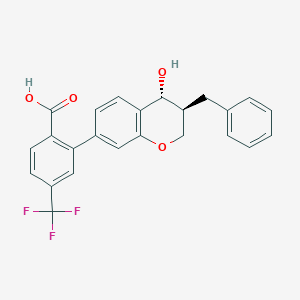 CP-195543 CP-195543 | C24H19F3O4 | 428.407 | 7 / 2 | 5.1 | No |
| 461299 | 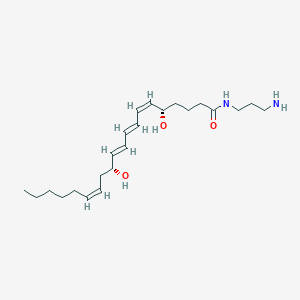 Leukotriene B4-3-aminopropylamide Leukotriene B4-3-aminopropylamide | C23H40N2O3 | 392.584 | 4 / 4 | 3.3 | Yes |
| 461378 | 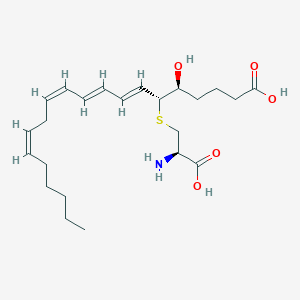 leukotriene E4 leukotriene E4 | C23H37NO5S | 439.611 | 7 / 4 | 2.1 | Yes |
| 250376 | 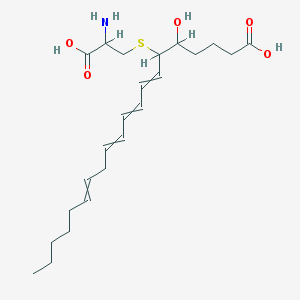 75715-89-8 75715-89-8 | C23H37NO5S | 439.611 | 7 / 4 | 2.1 | Yes |
| 461539 | 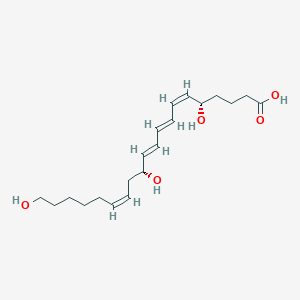 20-Hydroxy-leukotriene B4 20-Hydroxy-leukotriene B4 | C20H32O5 | 352.471 | 5 / 4 | 2.5 | Yes |
| 556519 | 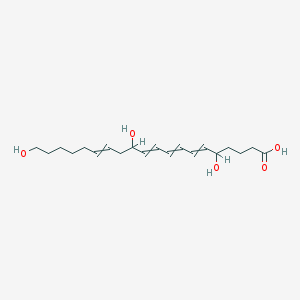 5,12,20-trihydroxy-6,8,10,14-eicosatetraenoic acid 5,12,20-trihydroxy-6,8,10,14-eicosatetraenoic acid | C20H32O5 | 352.471 | 5 / 4 | 2.5 | Yes |
| 272280 | 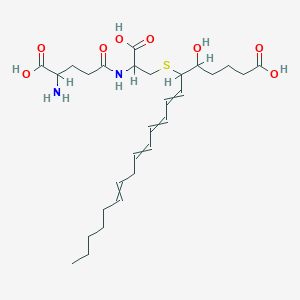 Leukotriene F4 Leukotriene F4 | C28H44N2O8S | 568.726 | 10 / 6 | 1.3 | No |
| 452678 | 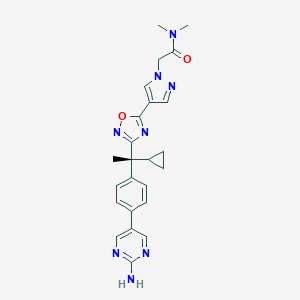 CHEMBL3417525 CHEMBL3417525 | C24H26N8O2 | 458.526 | 8 / 1 | 2.7 | Yes |
| 461741 | 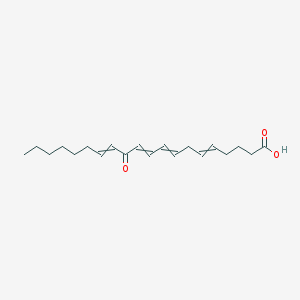 CID 91449379 CID 91449379 | C20H30O3 | 318.457 | 3 / 1 | 5.5 | No |
| 556636 | 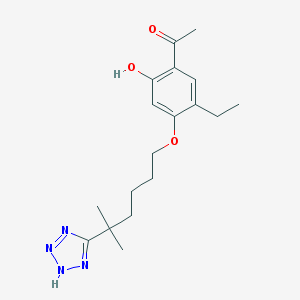 CHEMBL53132 CHEMBL53132 | C18H26N4O3 | 346.431 | 6 / 2 | 4.1 | Yes |
| 556705 | 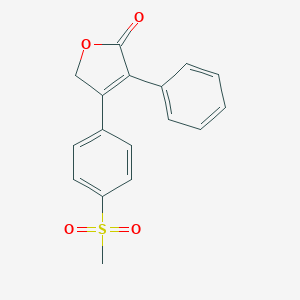 rofecoxib rofecoxib | C17H14O4S | 314.355 | 4 / 0 | 2.3 | Yes |
| 461892 | 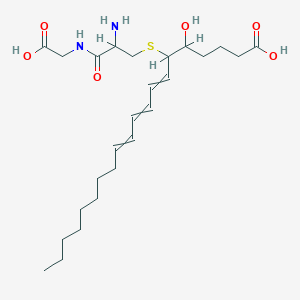 AC1MLYZ1 AC1MLYZ1 | C25H42N2O6S | 498.679 | 8 / 5 | 2.1 | Yes |
| 551768 | 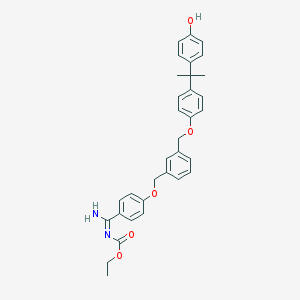 CID 60205731 CID 60205731 | C33H34N2O5 | 538.644 | 5 / 2 | 7.6 | No |
| 318022 | 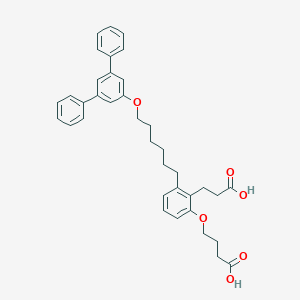 CHEMBL1099323 CHEMBL1099323 | C37H40O6 | 580.721 | 6 / 2 | 8.5 | No |
| 321515 | 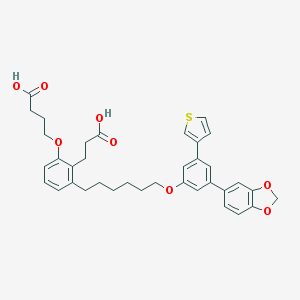 CHEMBL1098560 CHEMBL1098560 | C36H38O8S | 630.752 | 9 / 2 | 8.0 | No |
| 556784 | 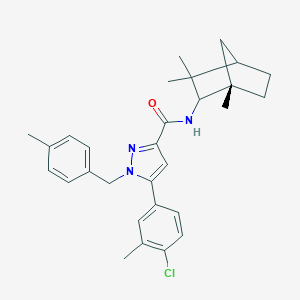 SCHEMBL1432281 SCHEMBL1432281 | C29H34ClN3O | 476.061 | 2 / 1 | 7.4 | No |
| 556804 | 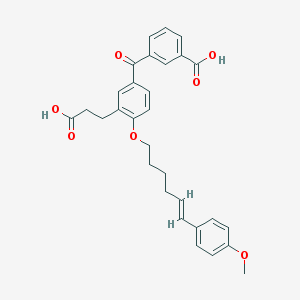 LY223982 LY223982 | C30H30O7 | 502.563 | 7 / 2 | 5.8 | No |
| 556827 | 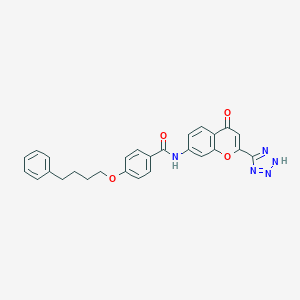 ONO-RS 411 ONO-RS 411 | C27H23N5O4 | 481.512 | 7 / 2 | 4.2 | Yes |
| 556935 | 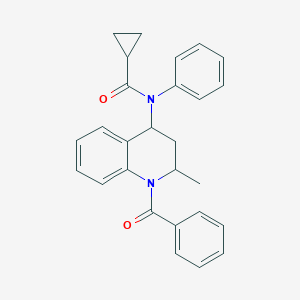 D03PYD D03PYD | C27H26N2O2 | 410.517 | 2 / 0 | 4.8 | Yes |
| 359908 | 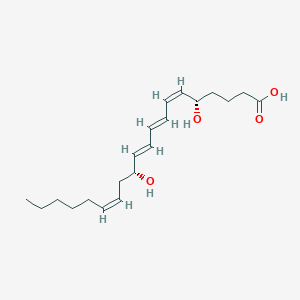 LEUKOTRIENE B4 LEUKOTRIENE B4 | C20H32O4 | 336.472 | 4 / 3 | 4.1 | Yes |
| 555075 | 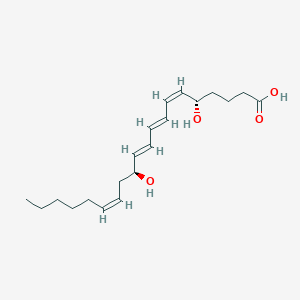 12-epi-LTB4 12-epi-LTB4 | C20H32O4 | 336.472 | 4 / 3 | 4.1 | Yes |
| 359913 | 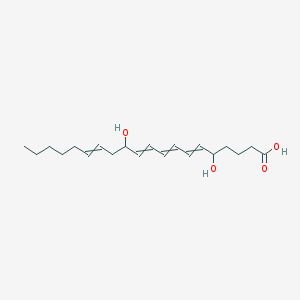 73151-67-4 73151-67-4 | C20H32O4 | 336.472 | 4 / 3 | 4.1 | Yes |
| 462314 | 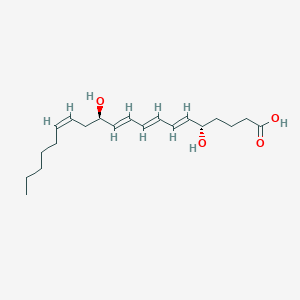 6-trans-Leukotriene B4 6-trans-Leukotriene B4 | C20H32O4 | 336.472 | 4 / 3 | 4.1 | Yes |
| 555083 | 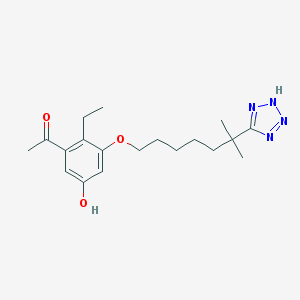 D0Y2IH D0Y2IH | C19H28N4O3 | 360.458 | 6 / 2 | 4.0 | Yes |
| 462367 | 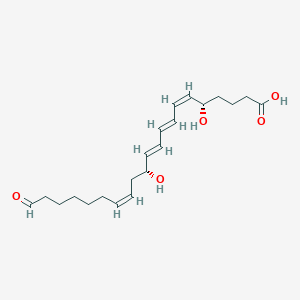 LTB4-20-carboxy LTB4-20-carboxy | C21H32O5 | 364.482 | 5 / 3 | 2.8 | Yes |
| 557025 | 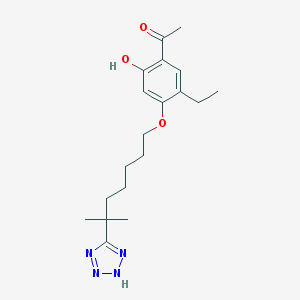 117690-79-6 117690-79-6 | C19H28N4O3 | 360.458 | 6 / 2 | 4.6 | Yes |
| 382009 | 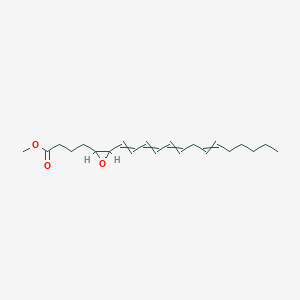 AC1N2PSN AC1N2PSN | C21H32O3 | 332.484 | 3 / 0 | 5.6 | No |
| 386444 | 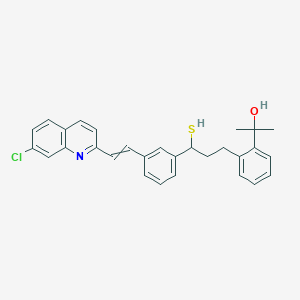 CID 77140346 CID 77140346 | C29H28ClNOS | 474.059 | 3 / 2 | 7.3 | No |
| 462690 | 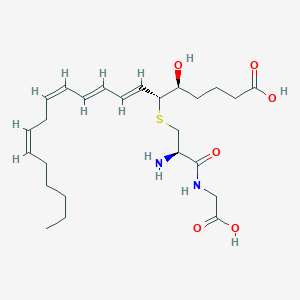 leukotriene D4 leukotriene D4 | C25H40N2O6S | 496.663 | 8 / 5 | 1.4 | Yes |
| 407982 | 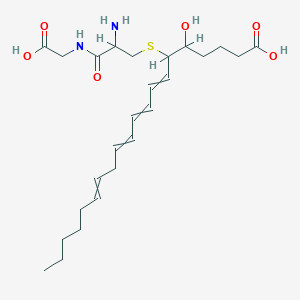 leukotriene D4 leukotriene D4 | C25H40N2O6S | 496.663 | 8 / 5 | 1.4 | Yes |
| 407998 | 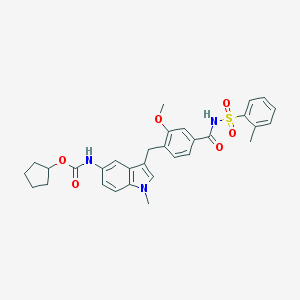 zafirlukast zafirlukast | C31H33N3O6S | 575.68 | 6 / 2 | 5.5 | No |
| 462740 | 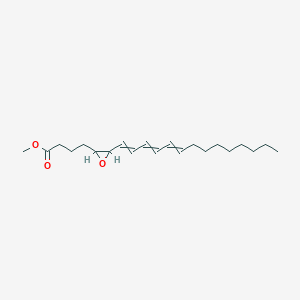 AC1N9SFB AC1N9SFB | C21H34O3 | 334.5 | 3 / 0 | 6.3 | No |
| 555430 | 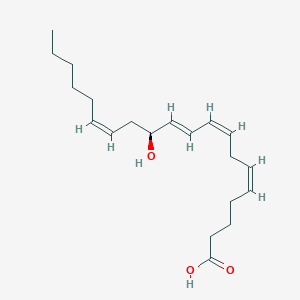 12(S)-HETE 12(S)-HETE | C20H32O3 | 320.473 | 3 / 2 | 5.2 | No |
| 557297 | 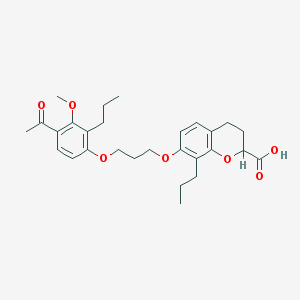 SC-41930 SC-41930 | C28H36O7 | 484.589 | 7 / 1 | 6.2 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218