

You can:
| Name | D(2) dopamine receptor |
|---|---|
| Species | Bos taurus (Bovine) |
| Gene | DRD2 |
| Synonym | Dopamine D2 receptor |
| Disease | N/A for non-human GPCRs |
| Length | 444 |
| Amino acid sequence | MDPLNLSWYDDDPESRNWSRPFNGSEGKADRPPYNYYAMLLTLLIFVIVFGNVLVCMAVSREKALQTTTNYLIVSLAVADLLVATLVMPWVVYLEVVGEWKFSRIHCDIFVTLDVMMCTASILNLCAISIDRYTAVAMPMLYNTRYSSKRRVTVMIAIVWVLSFTISCPMLFGLNNTDQNECIIANPAFVVYSSIVSFYVPFIVTLLVYIKIYIVLRRRRKRVNTKRSSRAFRANLKAPLKGNCTHPEDMKLCTVIMKSNGSFPVNRRRVEAARRAQELEMEMLSSTSPPERTRYSPIPPSHHQLTLPDPSHHGLHSTPDSPAKPEKNGHAKTVNPKIAKIFEIQSMPNGKTRTSLKTMSRRKLSQQKEKKATQMLAIVLGVFIICWLPFFITHILNIHCDCNIPPVLYSAFTWLGYVNSAVNPIIYTTFNIEFRKAFLKILHC |
| UniProt | P20288 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3998 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 42 | 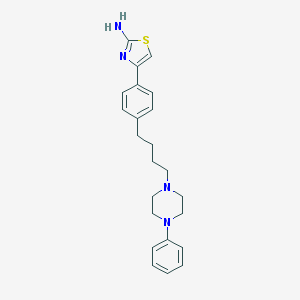 CHEMBL171316 CHEMBL171316 | C23H28N4S | 392.565 | 5 / 1 | 4.9 | Yes |
| 4224 | 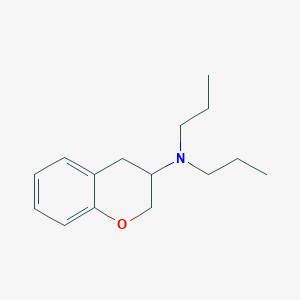 CHEMBL25984 CHEMBL25984 | C15H23NO | 233.355 | 2 / 0 | 3.8 | Yes |
| 555499 | 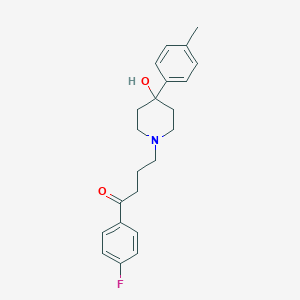 Moperone Moperone | C22H26FNO2 | 355.453 | 4 / 1 | 3.0 | Yes |
| 4977 | 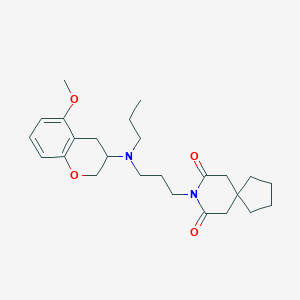 CHEMBL50188 CHEMBL50188 | C25H36N2O4 | 428.573 | 5 / 0 | 4.2 | Yes |
| 5536 | 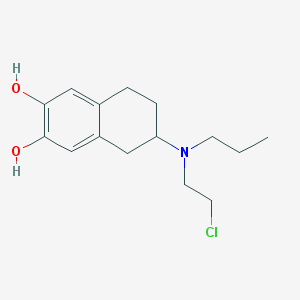 CHEMBL538235 CHEMBL538235 | C15H22ClNO2 | 283.796 | 3 / 2 | 3.5 | Yes |
| 5543 | 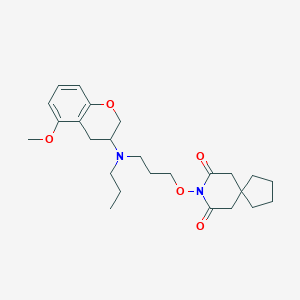 CHEMBL50722 CHEMBL50722 | C25H36N2O5 | 444.572 | 6 / 0 | 4.3 | Yes |
| 5549 | 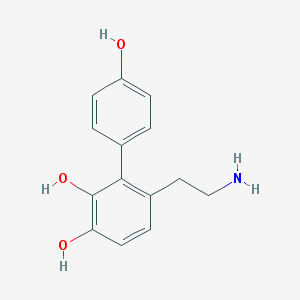 CHEMBL57470 CHEMBL57470 | C14H15NO3 | 245.278 | 4 / 4 | 1.8 | Yes |
| 5916 | 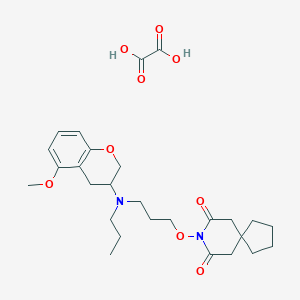 CHEMBL50722 CHEMBL50722 | C27H38N2O9 | 534.606 | 10 / 2 | N/A | No |
| 7287 | 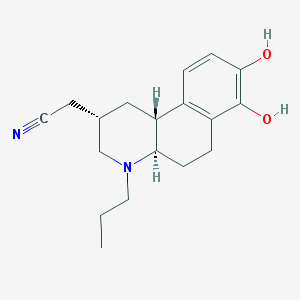 CHEMBL367343 CHEMBL367343 | C18H24N2O2 | 300.402 | 4 / 2 | 2.7 | Yes |
| 7311 | 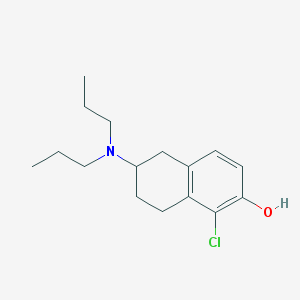 CHEMBL289557 CHEMBL289557 | C16H24ClNO | 281.824 | 2 / 1 | 4.7 | Yes |
| 9147 | 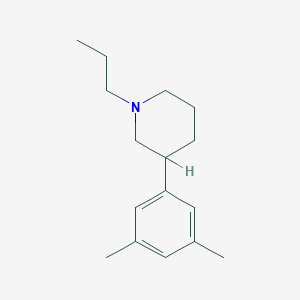 CHEMBL144114 CHEMBL144114 | C16H25N | 231.383 | 1 / 0 | 4.1 | Yes |
| 9598 | 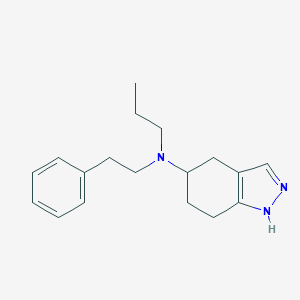 CHEMBL80437 CHEMBL80437 | C18H25N3 | 283.419 | 2 / 1 | 3.9 | Yes |
| 9766 | 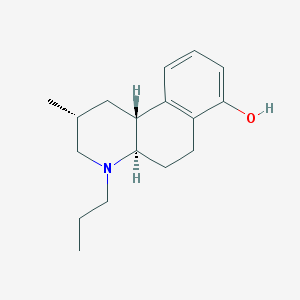 CHEMBL172205 CHEMBL172205 | C17H25NO | 259.393 | 2 / 1 | 3.9 | Yes |
| 10528 | 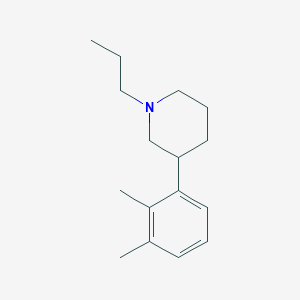 CHEMBL341825 CHEMBL341825 | C16H25N | 231.383 | 1 / 0 | 4.1 | Yes |
| 11217 | 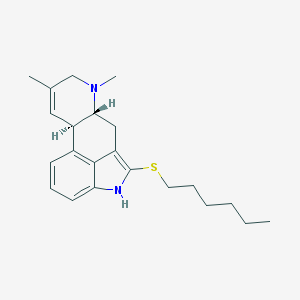 CHEMBL173042 CHEMBL173042 | C22H30N2S | 354.556 | 2 / 1 | 5.7 | No |
| 555526 | 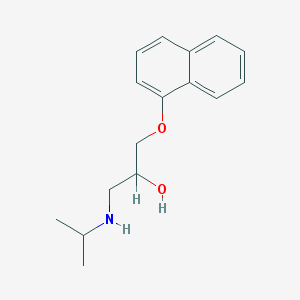 propranolol propranolol | C16H21NO2 | 259.349 | 3 / 2 | 3.0 | Yes |
| 11810 | 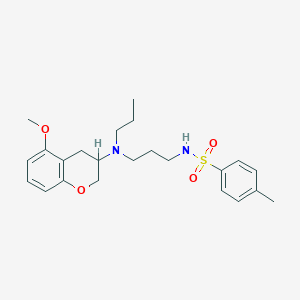 CHEMBL54266 CHEMBL54266 | C23H32N2O4S | 432.579 | 6 / 1 | 4.3 | Yes |
| 12432 | 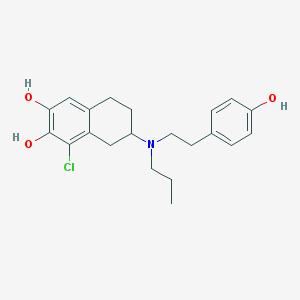 CHEMBL40949 CHEMBL40949 | C21H26ClNO3 | 375.893 | 4 / 3 | 5.1 | No |
| 12933 | 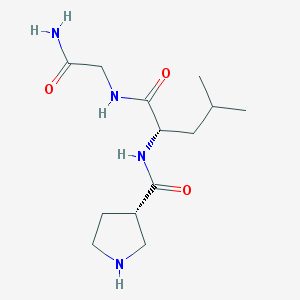 CHEMBL95208 CHEMBL95208 | C13H24N4O3 | 284.36 | 4 / 4 | -0.8 | Yes |
| 12934 | 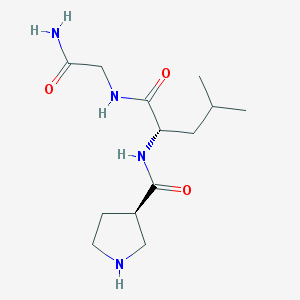 CHEMBL420258 CHEMBL420258 | C13H24N4O3 | 284.36 | 4 / 4 | -0.8 | Yes |
| 13620 | 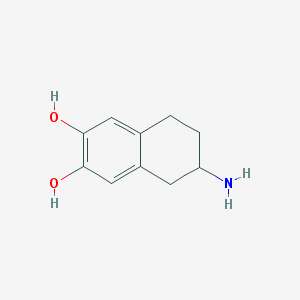 ADTN ADTN | C10H13NO2 | 179.219 | 3 / 3 | 1.4 | Yes |
| 13647 | 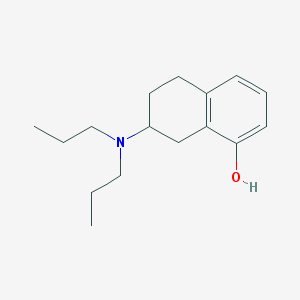 8-OH-Dpat 8-OH-Dpat | C16H25NO | 247.382 | 2 / 1 | 4.1 | Yes |
| 555551 | 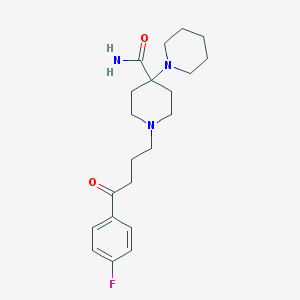 Pipamperone Pipamperone | C21H30FN3O2 | 375.488 | 5 / 1 | 2.0 | Yes |
| 17928 | 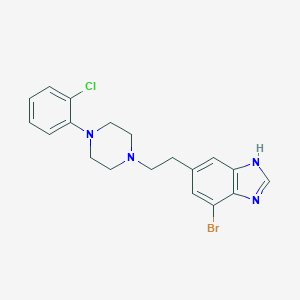 CHEMBL492042 CHEMBL492042 | C19H20BrClN4 | 419.751 | 3 / 1 | 4.7 | Yes |
| 555560 | 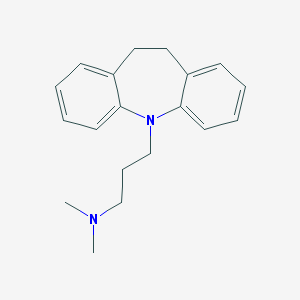 imipramine imipramine | C19H24N2 | 280.415 | 2 / 0 | 4.8 | Yes |
| 20298 | 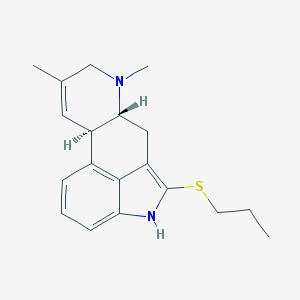 CHEMBL353848 CHEMBL353848 | C19H24N2S | 312.475 | 2 / 1 | 4.3 | Yes |
| 20618 | 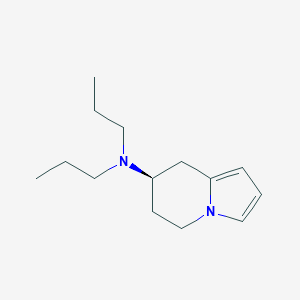 CHEMBL318924 CHEMBL318924 | C14H24N2 | 220.36 | 1 / 0 | 2.9 | Yes |
| 20760 | 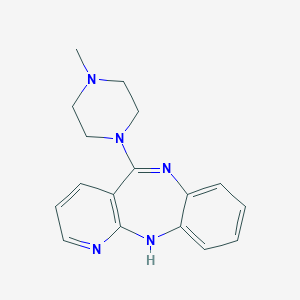 CHEMBL67355 CHEMBL67355 | C17H19N5 | 293.374 | 4 / 1 | 1.7 | Yes |
| 23377 | 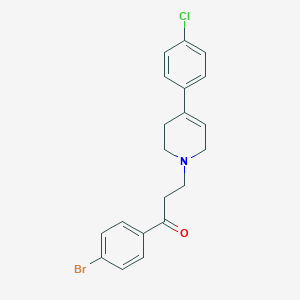 CHEMBL127067 CHEMBL127067 | C20H19BrClNO | 404.732 | 2 / 0 | 4.8 | Yes |
| 25086 | 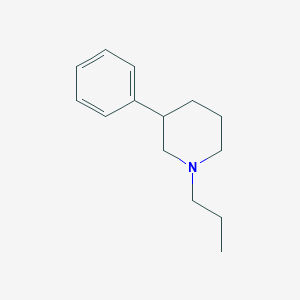 CHEMBL544711 CHEMBL544711 | C14H21N | 203.329 | 1 / 0 | 3.4 | Yes |
| 555576 | 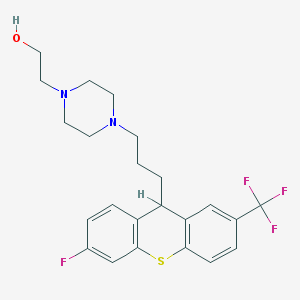 Teflutixol Teflutixol | C23H26F4N2OS | 454.528 | 8 / 1 | 4.6 | Yes |
| 27115 | 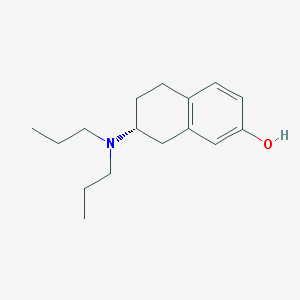 CHEMBL301559 CHEMBL301559 | C16H25NO | 247.382 | 2 / 1 | 4.1 | Yes |
| 29990 | 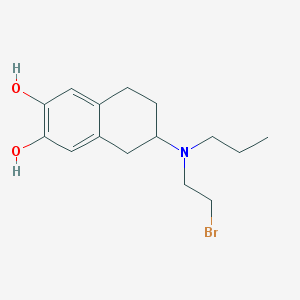 CHEMBL540272 CHEMBL540272 | C15H22BrNO2 | 328.25 | 3 / 2 | 3.6 | Yes |
| 30337 | 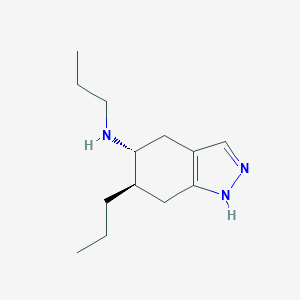 CHEMBL311360 CHEMBL311360 | C13H23N3 | 221.348 | 2 / 2 | 2.8 | Yes |
| 30395 | 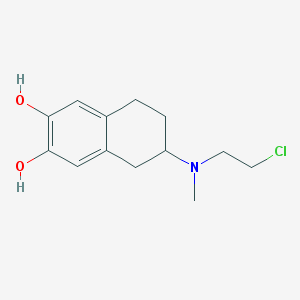 CHEMBL545203 CHEMBL545203 | C13H18ClNO2 | 255.742 | 3 / 2 | 2.6 | Yes |
| 32506 | 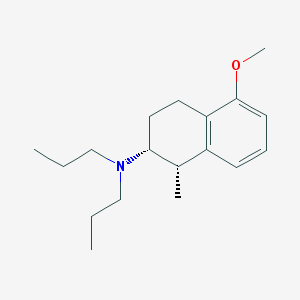 UNII-QQYOR9S587 UNII-QQYOR9S587 | C18H29NO | 275.436 | 2 / 0 | 4.8 | Yes |
| 33797 | 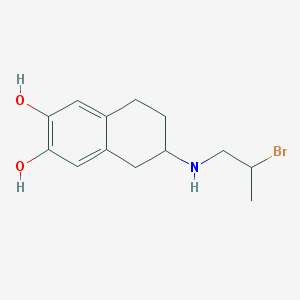 CHEMBL555369 CHEMBL555369 | C13H18BrNO2 | 300.196 | 3 / 3 | 2.7 | Yes |
| 35627 | 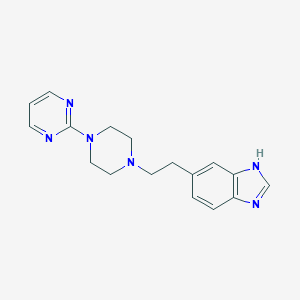 CHEMBL522639 CHEMBL522639 | C17H20N6 | 308.389 | 5 / 1 | 2.0 | Yes |
| 35938 | 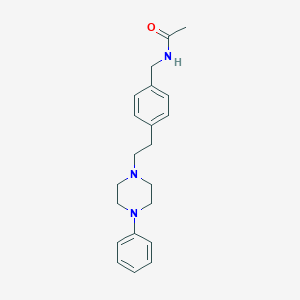 CHEMBL353678 CHEMBL353678 | C21H27N3O | 337.467 | 3 / 1 | 2.9 | Yes |
| 555623 | 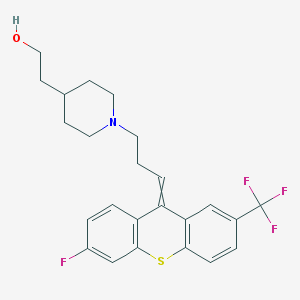 UNII-F52O5YF6CA UNII-F52O5YF6CA | C24H25F4NOS | 451.524 | 7 / 1 | 5.9 | No |
| 40943 | 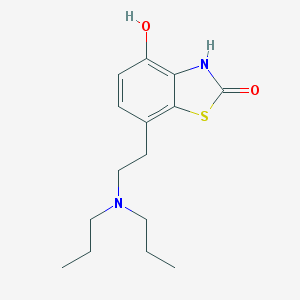 CHEMBL17231 CHEMBL17231 | C15H22N2O2S | 294.413 | 4 / 2 | 3.4 | Yes |
| 41057 | 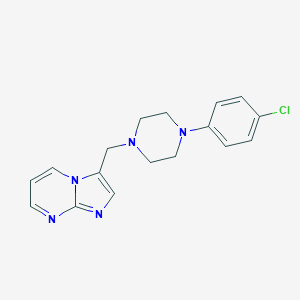 CHEMBL267853 CHEMBL267853 | C17H18ClN5 | 327.816 | 4 / 0 | 3.2 | Yes |
| 41311 | 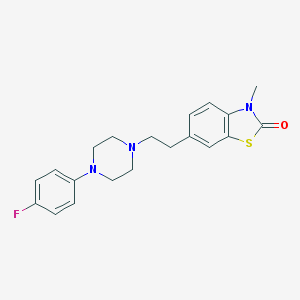 CHEMBL60318 CHEMBL60318 | C20H22FN3OS | 371.474 | 5 / 0 | 3.9 | Yes |
| 43789 | 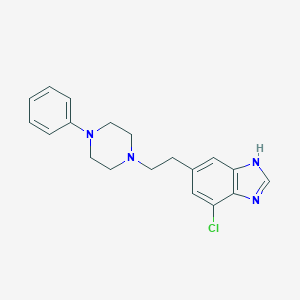 CHEMBL483864 CHEMBL483864 | C19H21ClN4 | 340.855 | 3 / 1 | 4.0 | Yes |
| 44471 | 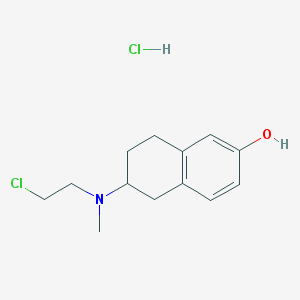 CHEMBL538745 CHEMBL538745 | C13H19Cl2NO | 276.201 | 2 / 2 | N/A | N/A |
| 44476 | 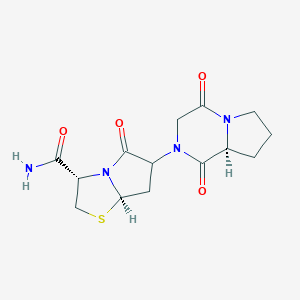 BDBM50060603 BDBM50060603 | C14H18N4O4S | 338.382 | 5 / 1 | -1.2 | Yes |
| 44477 | 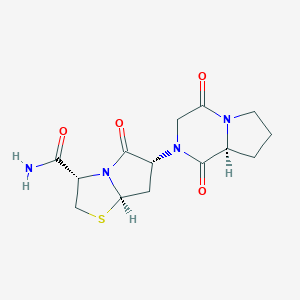 CHEMBL607228 CHEMBL607228 | C14H18N4O4S | 338.382 | 5 / 1 | -1.2 | Yes |
| 50419 | 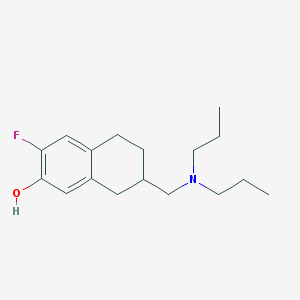 CHEMBL44255 CHEMBL44255 | C17H26FNO | 279.399 | 3 / 1 | 4.4 | Yes |
| 50875 | 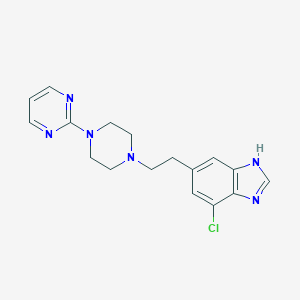 CHEMBL520019 CHEMBL520019 | C17H19ClN6 | 342.831 | 5 / 1 | 2.6 | Yes |
| 54612 | 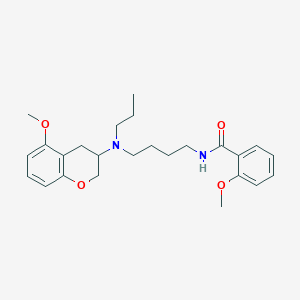 CHEMBL48925 CHEMBL48925 | C25H34N2O4 | 426.557 | 5 / 1 | 4.6 | Yes |
| 55119 | 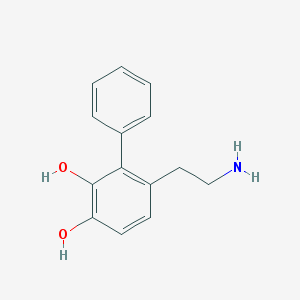 CHEMBL299511 CHEMBL299511 | C14H15NO2 | 229.279 | 3 / 3 | 2.2 | Yes |
| 56672 | 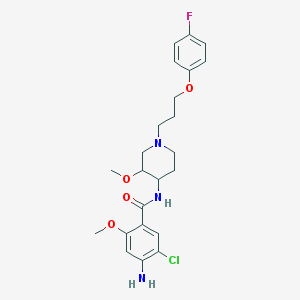 cisapride cisapride | C23H29ClFN3O4 | 465.95 | 7 / 2 | 3.4 | Yes |
| 56956 | 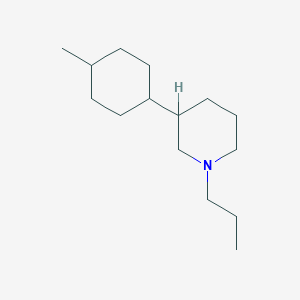 CHEMBL544961 CHEMBL544961 | C15H29N | 223.404 | 1 / 0 | 4.9 | Yes |
| 57047 | 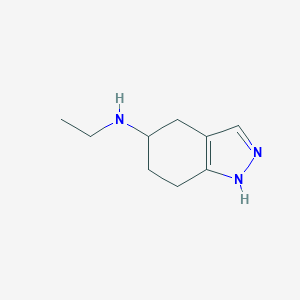 CHEMBL77421 CHEMBL77421 | C9H15N3 | 165.24 | 2 / 2 | 0.9 | Yes |
| 57055 | 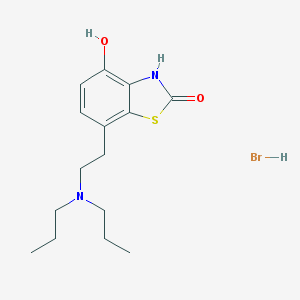 CHEMBL543212 CHEMBL543212 | C15H23BrN2O2S | 375.325 | 4 / 3 | N/A | N/A |
| 57379 | 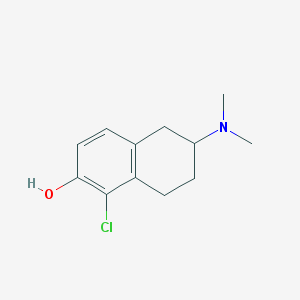 CHEMBL44419 CHEMBL44419 | C12H16ClNO | 225.716 | 2 / 1 | 3.0 | Yes |
| 57405 | 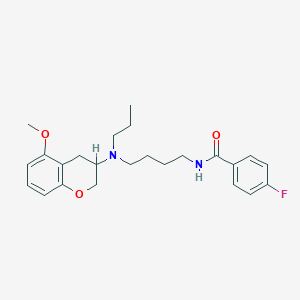 CHEMBL298534 CHEMBL298534 | C24H31FN2O3 | 414.521 | 5 / 1 | 4.7 | Yes |
| 59261 | 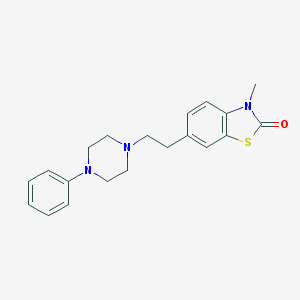 CHEMBL64167 CHEMBL64167 | C20H23N3OS | 353.484 | 4 / 0 | 3.8 | Yes |
| 555716 | 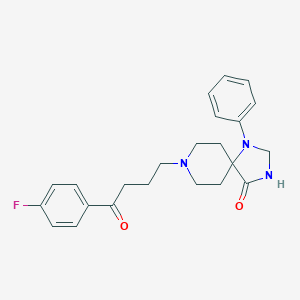 spiperone spiperone | C23H26FN3O2 | 395.478 | 5 / 1 | 3.0 | Yes |
| 62320 | 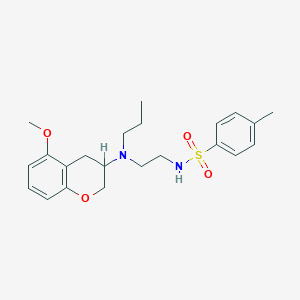 CHEMBL51888 CHEMBL51888 | C22H30N2O4S | 418.552 | 6 / 1 | 3.9 | Yes |
| 62509 | 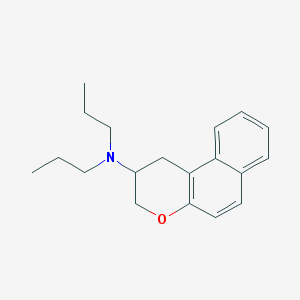 CHEMBL52438 CHEMBL52438 | C19H25NO | 283.415 | 2 / 0 | 5.1 | No |
| 63057 | 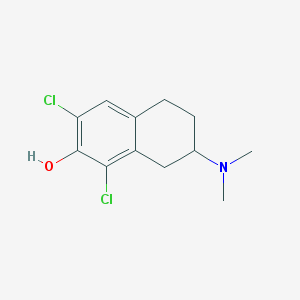 CHEMBL418252 CHEMBL418252 | C12H15Cl2NO | 260.158 | 2 / 1 | 3.6 | Yes |
| 63075 | 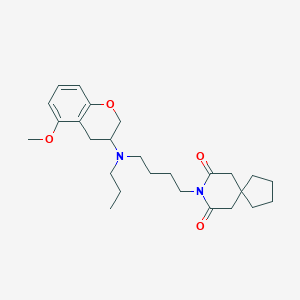 CHEMBL11592 CHEMBL11592 | C26H38N2O4 | 442.6 | 5 / 0 | 4.6 | Yes |
| 63281 | 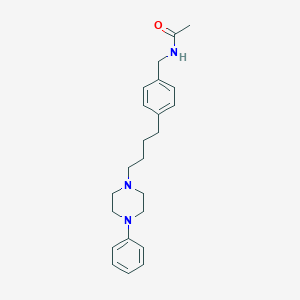 CHEMBL169702 CHEMBL169702 | C23H31N3O | 365.521 | 3 / 1 | 3.6 | Yes |
| 64670 | 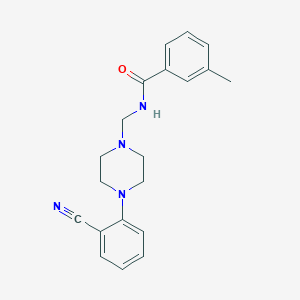 PD-168077 PD-168077 | C20H22N4O | 334.423 | 4 / 1 | 3.0 | Yes |
| 64704 | 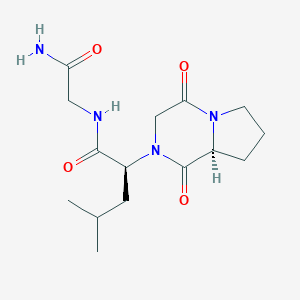 CHEMBL332740 CHEMBL332740 | C15H24N4O4 | 324.381 | 4 / 2 | -0.4 | Yes |
| 64764 | 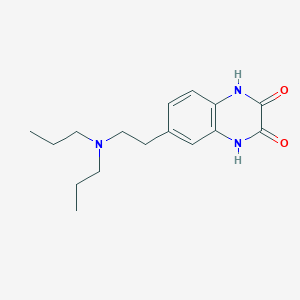 CHEMBL338119 CHEMBL338119 | C16H23N3O2 | 289.379 | 3 / 2 | 2.5 | Yes |
| 65589 | 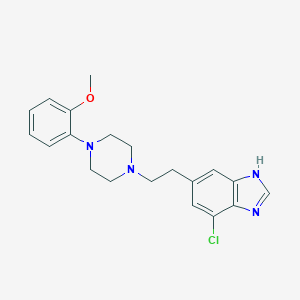 CHEMBL510055 CHEMBL510055 | C20H23ClN4O | 370.881 | 4 / 1 | 4.0 | Yes |
| 66338 | 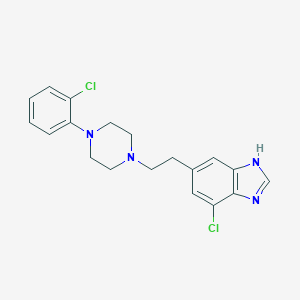 CHEMBL483247 CHEMBL483247 | C19H20Cl2N4 | 375.297 | 3 / 1 | 4.6 | Yes |
| 66640 | 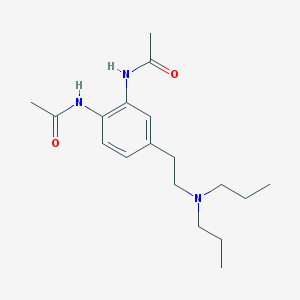 CHEMBL131186 CHEMBL131186 | C18H29N3O2 | 319.449 | 3 / 2 | 2.4 | Yes |
| 66729 | 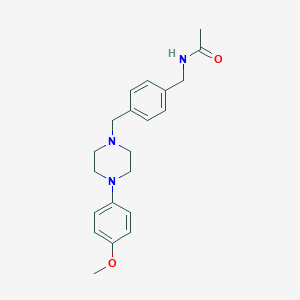 CHEMBL171115 CHEMBL171115 | C21H27N3O2 | 353.466 | 4 / 1 | 2.4 | Yes |
| 67253 | 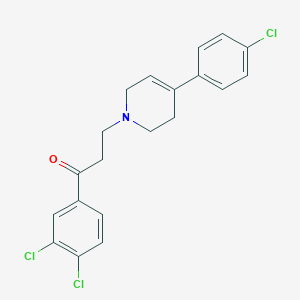 CHEMBL340541 CHEMBL340541 | C20H18Cl3NO | 394.72 | 2 / 0 | 5.3 | No |
| 67353 | 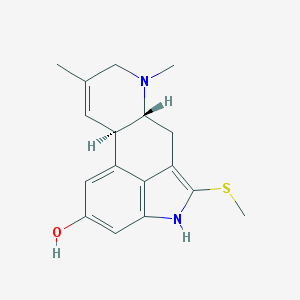 CHEMBL170356 CHEMBL170356 | C17H20N2OS | 300.42 | 3 / 2 | 3.0 | Yes |
| 67843 | 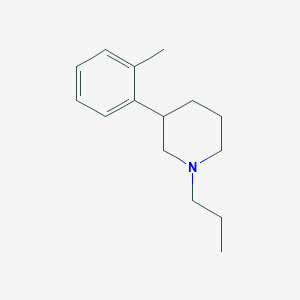 CHEMBL553258 CHEMBL553258 | C15H23N | 217.356 | 1 / 0 | 3.8 | Yes |
| 68310 | 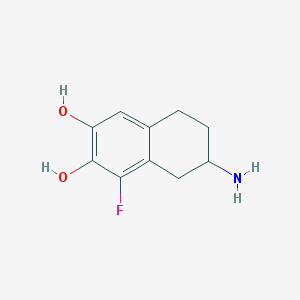 CHEMBL432676 CHEMBL432676 | C10H12FNO2 | 197.209 | 4 / 3 | 1.1 | Yes |
| 68933 | 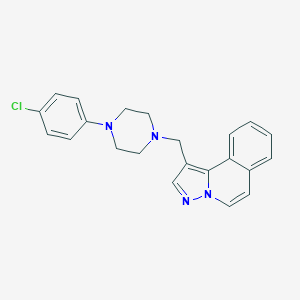 CHEMBL268558 CHEMBL268558 | C22H21ClN4 | 376.888 | 3 / 0 | 4.4 | Yes |
| 68948 | 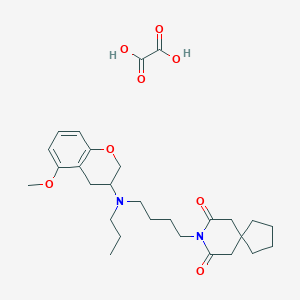 CHEMBL301060 CHEMBL301060 | C28H40N2O8 | 532.634 | 9 / 2 | N/A | No |
| 71268 | 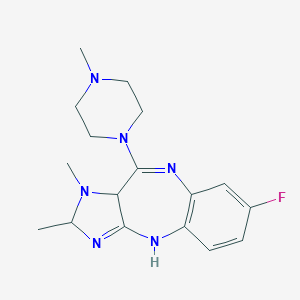 BDBM50017525 BDBM50017525 | C17H23FN6 | 330.411 | 5 / 1 | 0.7 | Yes |
| 72741 | 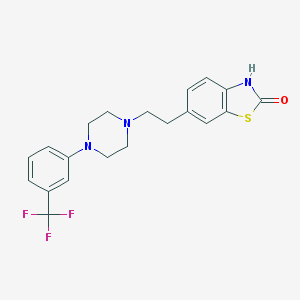 CHEMBL59741 CHEMBL59741 | C20H20F3N3OS | 407.455 | 7 / 1 | 4.5 | Yes |
| 73740 | 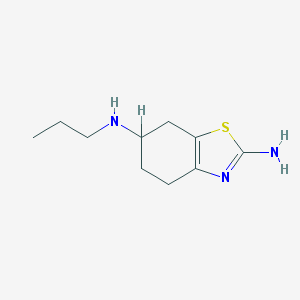 FASDKYOPVNHBLU-UHFFFAOYSA-N FASDKYOPVNHBLU-UHFFFAOYSA-N | C10H17N3S | 211.327 | 4 / 2 | 1.9 | Yes |
| 73744 | 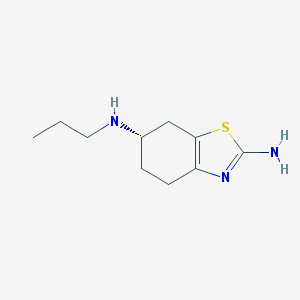 pramipexole pramipexole | C10H17N3S | 211.327 | 4 / 2 | 1.9 | Yes |
| 74350 | 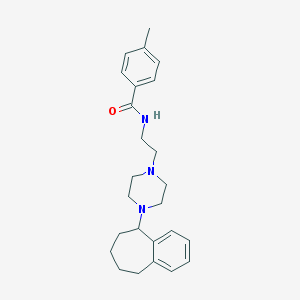 CHEMBL176777 CHEMBL176777 | C25H33N3O | 391.559 | 3 / 1 | 4.2 | Yes |
| 75809 | 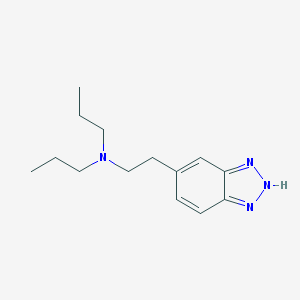 CHEMBL131120 CHEMBL131120 | C14H22N4 | 246.358 | 3 / 1 | 3.1 | Yes |
| 555762 | 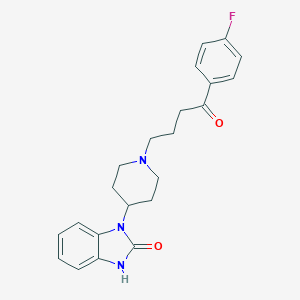 BENPERIDOL BENPERIDOL | C22H24FN3O2 | 381.451 | 4 / 1 | 3.7 | Yes |
| 76585 | 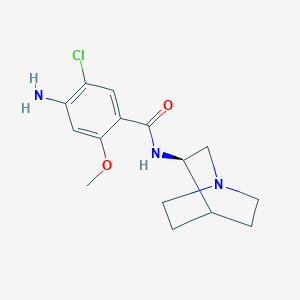 (R)-zacopride (R)-zacopride | C15H20ClN3O2 | 309.794 | 4 / 2 | 1.7 | Yes |
| 77554 | 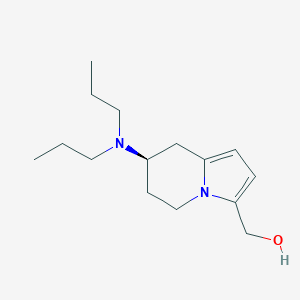 CHEMBL99291 CHEMBL99291 | C15H26N2O | 250.386 | 2 / 1 | 2.1 | Yes |
| 77558 | 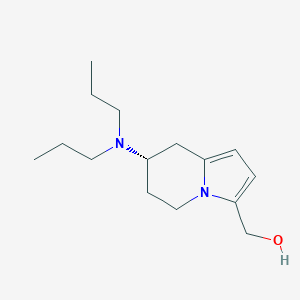 CHEMBL95230 CHEMBL95230 | C15H26N2O | 250.386 | 2 / 1 | 2.1 | Yes |
| 77922 | 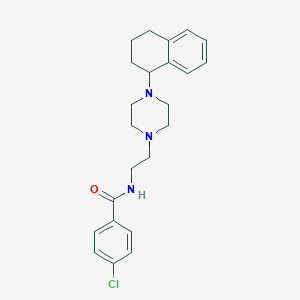 CHEMBL177031 CHEMBL177031 | C23H28ClN3O | 397.947 | 3 / 1 | 4.0 | Yes |
| 82457 | 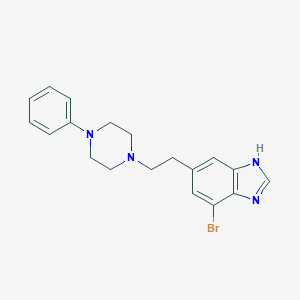 CHEMBL485499 CHEMBL485499 | C19H21BrN4 | 385.309 | 3 / 1 | 4.1 | Yes |
| 83677 | 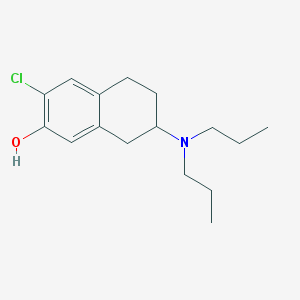 CHEMBL295196 CHEMBL295196 | C16H24ClNO | 281.824 | 2 / 1 | 4.7 | Yes |
| 84878 | 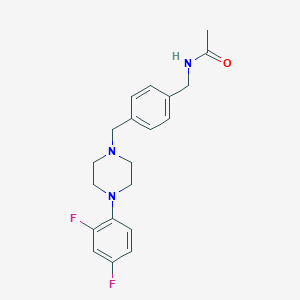 CHEMBL352576 CHEMBL352576 | C20H23F2N3O | 359.421 | 5 / 1 | 2.7 | Yes |
| 87065 | 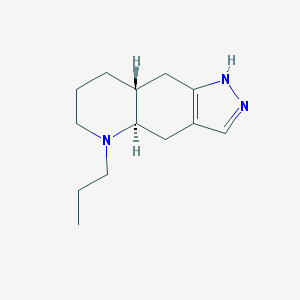 QUINPIROLE QUINPIROLE | C13H21N3 | 219.332 | 2 / 1 | 2.3 | Yes |
| 88085 | 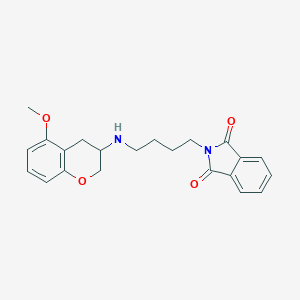 CHEMBL296395 CHEMBL296395 | C22H24N2O4 | 380.444 | 5 / 1 | 2.9 | Yes |
| 88273 | 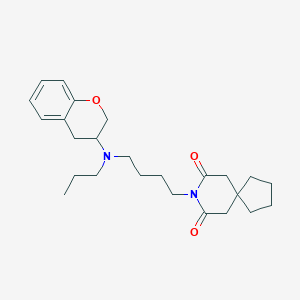 CHEMBL50993 CHEMBL50993 | C25H36N2O3 | 412.574 | 4 / 0 | 4.6 | Yes |
| 88559 | 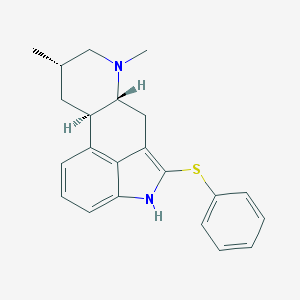 CHEMBL354362 CHEMBL354362 | C22H24N2S | 348.508 | 2 / 1 | 5.6 | No |
| 88560 | 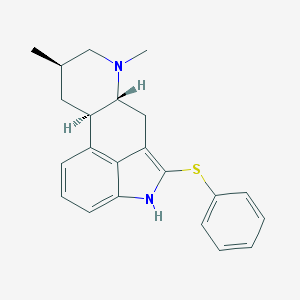 CHEMBL435263 CHEMBL435263 | C22H24N2S | 348.508 | 2 / 1 | 5.6 | No |
| 89902 | 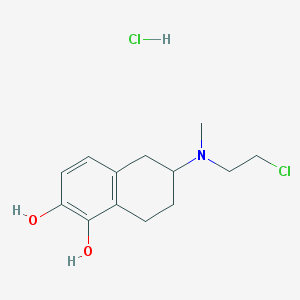 CHEMBL544038 CHEMBL544038 | C13H19Cl2NO2 | 292.2 | 3 / 3 | N/A | N/A |
| 90087 | 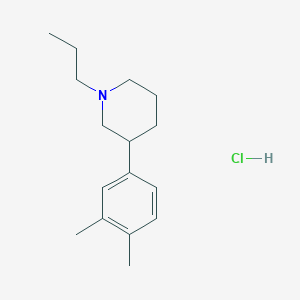 3-(3,4-Dimethylphenyl)-1-propyl-piperidine hydrochloride 3-(3,4-Dimethylphenyl)-1-propyl-piperidine hydrochloride | C16H26ClN | 267.841 | 1 / 1 | N/A | N/A |
| 92987 | 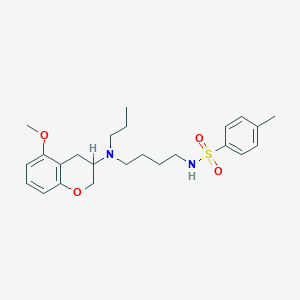 CHEMBL300735 CHEMBL300735 | C24H34N2O4S | 446.606 | 6 / 1 | 4.6 | Yes |
| 555818 | 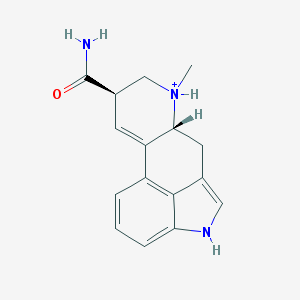 D-Lysergic acid amide D-Lysergic acid amide | C16H18N3O+ | 268.34 | 1 / 3 | 1.6 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218