

You can:
| Name | Nociceptin receptor |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Oprl1 |
| Synonym | NOP receptor NOP-r NOPr OP4 ORGC [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 367 |
| Amino acid sequence | MESLFPAPFWEVLYGSHFQGNLSLLNETVPHHLLLNASHSAFLPLGLKVTIVGLYLAVCIGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLLTLPFQGTDILLGFWPFGNALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPIRALDVRTSSKAQAVNVAIWALASVVGVPVAIMGSAQVEDEEIECLVEIPAPQDYWGPVFAICIFLFSFIIPVLIISVCYSLMIRRLRGVRLLSGSREKDRNLRRITRLVLVVVAVFVGCWTPVQVFVLVQGLGVQPGSETAVAILRFCTALGYVNSCLNPILYAFLDENFKACFRKFCCASALHREMQVSDRVRSIAKDVGLGCKTSETVPRPA |
| UniProt | P35377 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3621 |
| IUPHAR | 320 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 555484 | 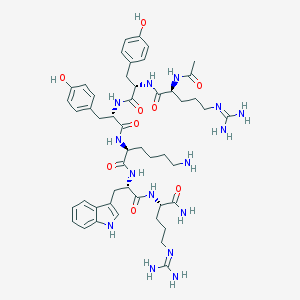 Ac-RYYKWR-NH2 Ac-RYYKWR-NH2 | C49H69N15O9 | 1012.19 | 12 / 15 | -1.3 | No |
| 555502 | 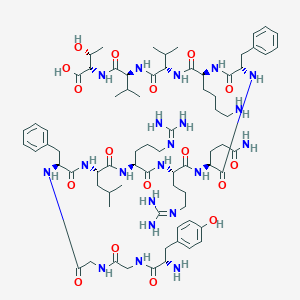 Dynorphin B Dynorphin B | C74H115N21O17 | 1570.86 | 21 / 22 | -3.6 | No |
| 555510 | 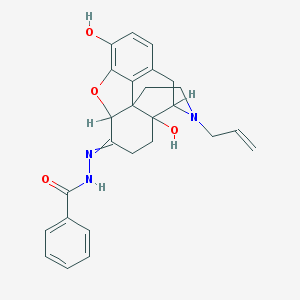 AC1L1I54 AC1L1I54 | C26H27N3O4 | 445.519 | 6 / 3 | 3.5 | Yes |
| 20202 | 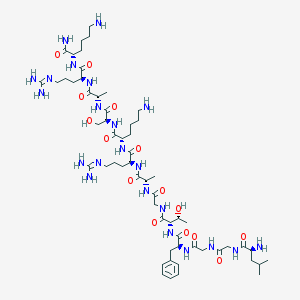 CHEMBL412479 CHEMBL412479 | C58H102N22O15 | 1347.59 | 20 / 22 | -7.2 | No |
| 555562 | 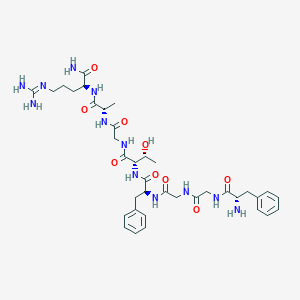 BDBM85820 BDBM85820 | C37H54N12O9 | 810.914 | 11 / 12 | -3.0 | No |
| 22788 | 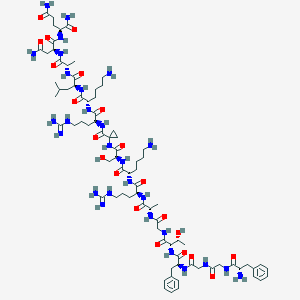 CHEMBL396972 CHEMBL396972 | C80H130N28O21 | 1820.09 | 26 / 30 | -8.1 | No |
| 25218 | 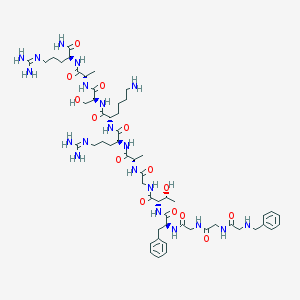 CHEMBL428167 CHEMBL428167 | C55H88N20O14 | 1253.44 | 18 / 20 | -6.5 | No |
| 555597 | 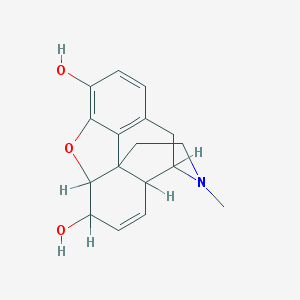 Morphinan-3,6.alpha.-diol, 7,8-didehydro-4,5.alpha.-epoxy-17-methyl- Morphinan-3,6.alpha.-diol, 7,8-didehydro-4,5.alpha.-epoxy-17-methyl- | C17H19NO3 | 285.343 | 4 / 2 | 0.8 | Yes |
| 36383 | 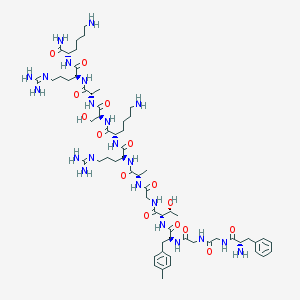 CHEMBL262224 CHEMBL262224 | C62H102N22O15 | 1395.64 | 20 / 22 | -6.6 | No |
| 48792 | 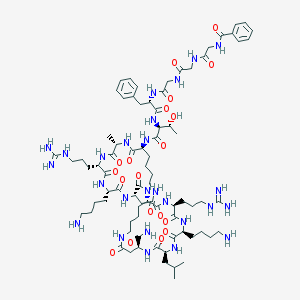 CID 53316770 CID 53316770 | C79H126N26O19 | 1744.04 | 23 / 27 | -4.8 | No |
| 52229 | 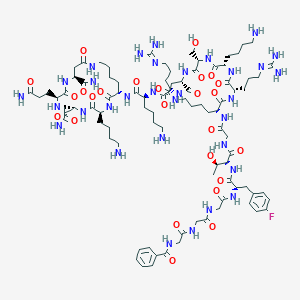 BDBM50333103 BDBM50333103 | C90H143FN32O25 | 2092.33 | 31 / 32 | -11.9 | No |
| 52495 | 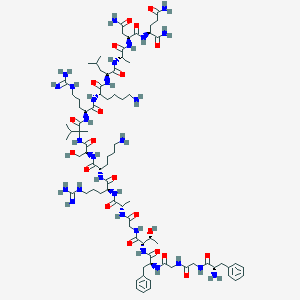 CHEMBL390658 CHEMBL390658 | C82H136N28O21 | 1850.17 | 26 / 30 | -7.4 | No |
| 61754 | 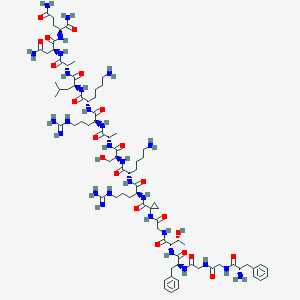 CHEMBL394775 CHEMBL394775 | C80H130N28O21 | 1820.09 | 26 / 30 | -8.6 | No |
| 555732 | 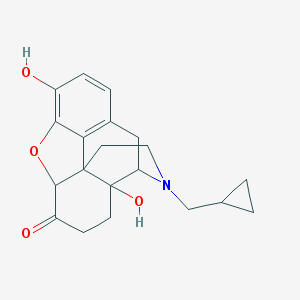 DQCKKXVULJGBQN-UHFFFAOYSA-N DQCKKXVULJGBQN-UHFFFAOYSA-N | C20H23NO4 | 341.407 | 5 / 2 | 1.9 | Yes |
| 69464 | 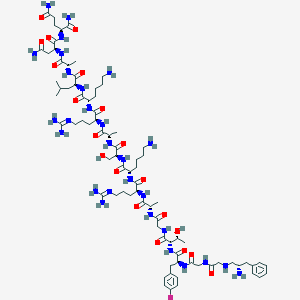 CHEMBL263588 CHEMBL263588 | C79H131FN28O20 | 1812.09 | 27 / 28 | -9.0 | No |
| 71073 | 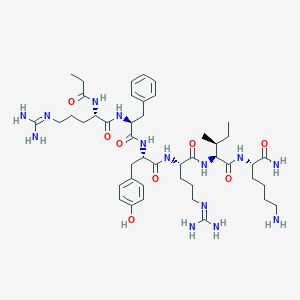 CHEMBL403807 CHEMBL403807 | C45H72N14O8 | 937.161 | 11 / 13 | -0.8 | No |
| 82690 |  CID 100916438 CID 100916438 | C79H130N28O21 | 1808.08 | 45 / 33 | 2.1 | No |
| 86603 |  CHEMBL541911 CHEMBL541911 | C82H137ClN28O21 | 1886.62 | 26 / 31 | N/A | No |
| 90700 |  CHEMBL407461 CHEMBL407461 | C61H102N22O15 | 1383.63 | 21 / 23 | -7.1 | No |
| 93251 |  CHEMBL405648 CHEMBL405648 | C61H99ClN22O15 | 1416.05 | 20 / 22 | -6.3 | No |
| 93356 | 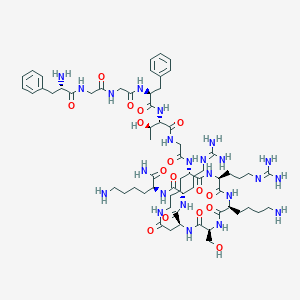 CHEMBL1631925 CHEMBL1631925 | C65H105N23O16 | 1464.7 | 21 / 23 | -7.1 | No |
| 555807 | 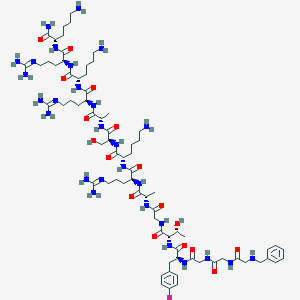 OFQ/N UFP-102 OFQ/N UFP-102 | C73H123FN28O17 | 1683.96 | 25 / 27 | -8.8 | No |
| 93499 | 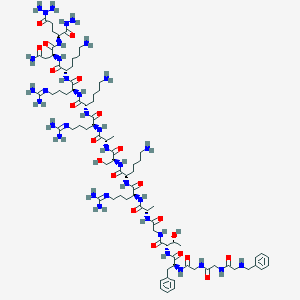 CHEMBL410145 CHEMBL410145 | C82H141N35O21 | 1953.25 | 31 / 33 | -13.3 | No |
| 99216 | 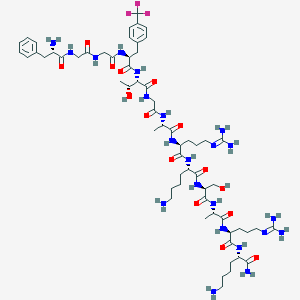 CHEMBL265544 CHEMBL265544 | C62H99F3N22O15 | 1449.61 | 23 / 22 | -6.1 | No |
| 555855 | 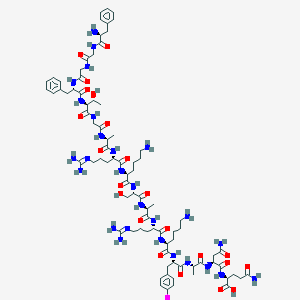 OFQ/N,Iodo[Tyr14] OFQ/N,Iodo[Tyr14] | C82H126IN27O22 | 1968.98 | 27 / 28 | -10.1 | No |
| 106290 | 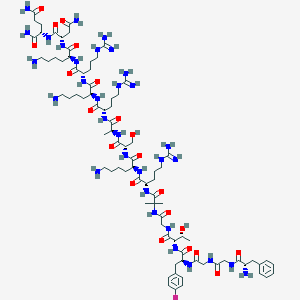 UFP-112 UFP-112 | C83H139FN32O21 | 1940.23 | 29 / 34 | -10.7 | No |
| 107028 | 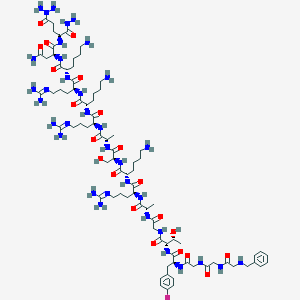 CHEMBL411649 CHEMBL411649 | C82H140FN35O21 | 1971.24 | 32 / 33 | -13.2 | No |
| 555888 | 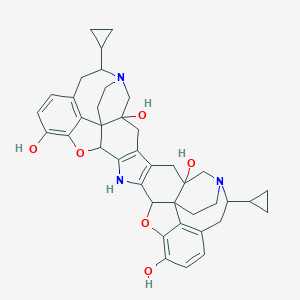 AC1MHXH5 AC1MHXH5 | C40H43N3O6 | 661.799 | 8 / 5 | 3.2 | No |
| 119428 | 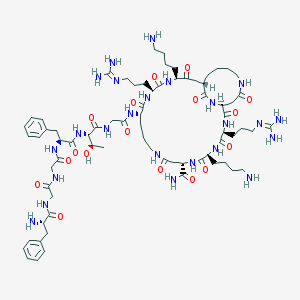 CHEMBL1631924 CHEMBL1631924 | C73H116N24O17 | 1601.88 | 22 / 23 | -7.1 | No |
| 555958 | 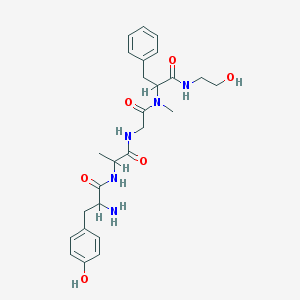 AC1NB6ST AC1NB6ST | C26H35N5O6 | 513.595 | 7 / 6 | 0.2 | No |
| 126215 | 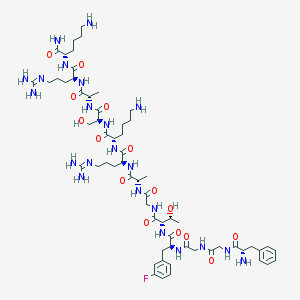 CHEMBL412537 CHEMBL412537 | C61H99FN22O15 | 1399.6 | 21 / 22 | -6.9 | No |
| 126429 | 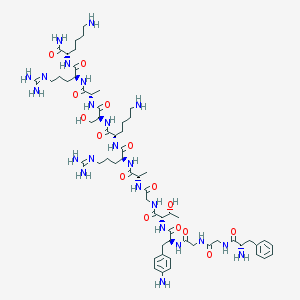 CHEMBL262928 CHEMBL262928 | C61H101N23O15 | 1396.62 | 21 / 23 | -7.7 | No |
| 142473 | 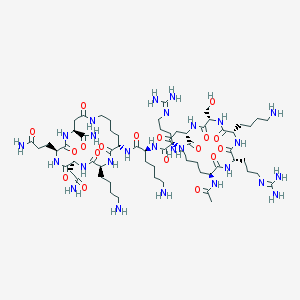 CHEMBL1631933 CHEMBL1631933 | C64H114N26O18 | 1535.78 | 23 / 25 | -11.5 | No |
| 149023 | 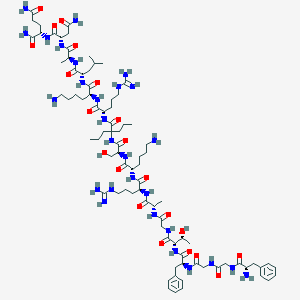 CHEMBL394588 CHEMBL394588 | C84H140N28O21 | 1878.22 | 26 / 30 | -6.6 | No |
| 150650 | 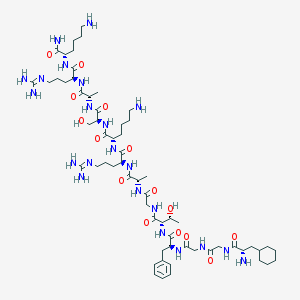 CHEMBL406905 CHEMBL406905 | C61H106N22O15 | 1387.66 | 20 / 22 | -5.8 | No |
| 152018 | 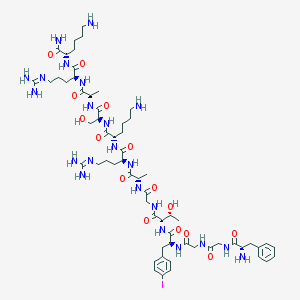 CHEMBL410167 CHEMBL410167 | C61H99IN22O15 | 1507.51 | 20 / 22 | -6.3 | No |
| 556055 | 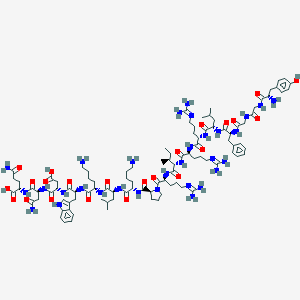 H-Tyr-Gly-Gly-Phe-Leu-Arg-Arg-Ile-Arg-Pro-Lys-Leu-Lys-Trp-Asp-Asn-Gln-OH H-Tyr-Gly-Gly-Phe-Leu-Arg-Arg-Ile-Arg-Pro-Lys-Leu-Lys-Trp-Asp-Asn-Gln-OH | C99H155N31O23 | 2147.52 | 29 / 30 | -8.9 | No |
| 170954 | 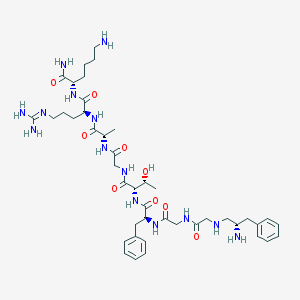 CHEMBL317625 CHEMBL317625 | C43H68N14O9 | 925.106 | 13 / 14 | -3.2 | No |
| 180435 | 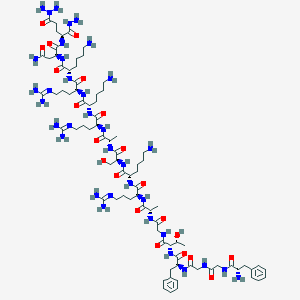 CHEMBL410653 CHEMBL410653 | C82H141N35O21 | 1953.25 | 31 / 33 | -13.4 | No |
| 556207 | 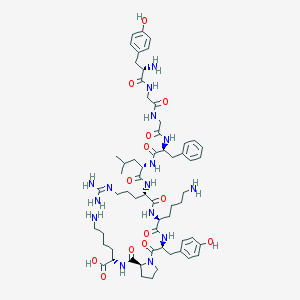 alpha-Neoendorphin alpha-Neoendorphin | C60H89N15O13 | 1228.46 | 17 / 16 | -2.1 | No |
| 183238 | 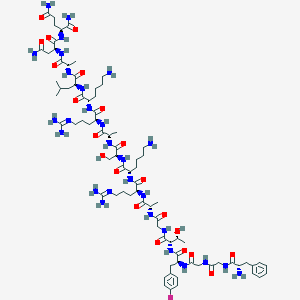 CHEMBL264846 CHEMBL264846 | C79H129FN28O21 | 1826.07 | 27 / 28 | -9.2 | No |
| 183733 | 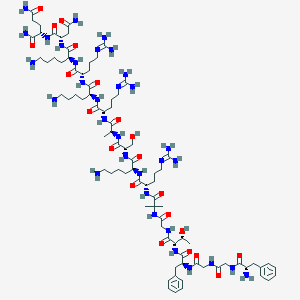 BDBM50411363 BDBM50411363 | C83H140N32O21 | 1922.24 | 28 / 31 | -11.9 | No |
| 554243 | 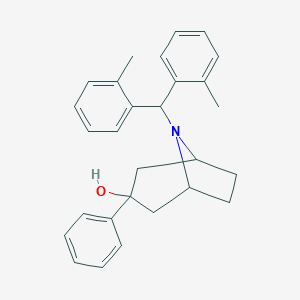 SCH 221510 SCH 221510 | C28H31NO | 397.562 | 2 / 1 | 6.0 | No |
| 200764 | 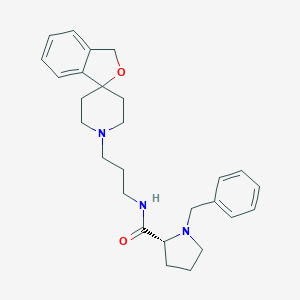 475150-69-7 475150-69-7 | C27H35N3O2 | 433.596 | 4 / 1 | 3.3 | Yes |
| 556284 | 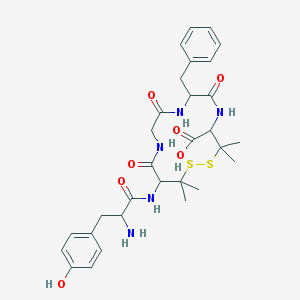 88373-73-3 88373-73-3 | C30H39N5O7S2 | 645.79 | 10 / 7 | -0.6 | No |
| 203601 | 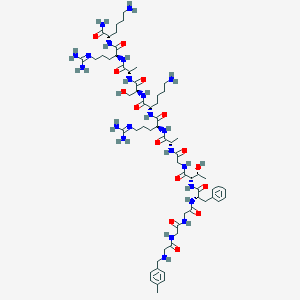 CHEMBL411168 CHEMBL411168 | C62H102N22O15 | 1395.64 | 20 / 22 | -6.6 | No |
| 210372 | 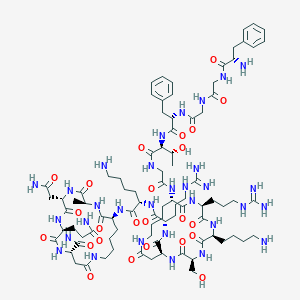 CID 49850694 CID 49850694 | C87H139N31O24 | 2003.26 | 29 / 33 | -10.9 | No |
| 215309 | 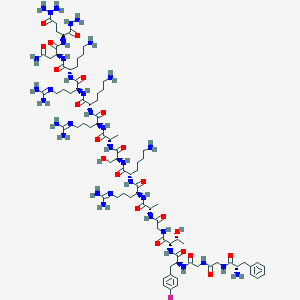 CHEMBL414782 CHEMBL414782 | C82H140FN35O21 | 1971.24 | 32 / 33 | -13.3 | No |
| 215323 |  CHEMBL232245 CHEMBL232245 | C82H136N28O21 | 1850.17 | 26 / 30 | -7.4 | No |
| 215445 |  CHEMBL414963 CHEMBL414963 | C62H104N22O14 | 1381.65 | 20 / 22 | -6.3 | No |
| 219902 |  CHEMBL438398 CHEMBL438398 | C82H136N28O21 | 1850.17 | 26 / 30 | -7.3 | No |
| 223861 |  CHEMBL412606 CHEMBL412606 | C61H100N22O16 | 1397.61 | 21 / 23 | -7.3 | No |
| 224036 |  CHEMBL556556 CHEMBL556556 | C82H137ClN28O21 | 1886.62 | 26 / 31 | N/A | No |
| 226737 |  [Nphe1]nociceptin(1-13)NH2 [Nphe1]nociceptin(1-13)NH2 | C61H100N22O15 | 1381.61 | 20 / 22 | -7.0 | No |
| 231305 |  CHEMBL427699 CHEMBL427699 | C81H134N28O21 | 1836.14 | 26 / 30 | -7.9 | No |
| 556389 |  CID 102330627 CID 102330627 | C82H127N27O23 | 1859.08 | 46 / 33 | -0.4 | No |
| 236895 | 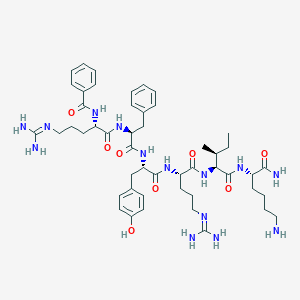 CHEMBL257921 CHEMBL257921 | C49H72N14O8 | 985.205 | 11 / 13 | 0.4 | No |
| 240684 | 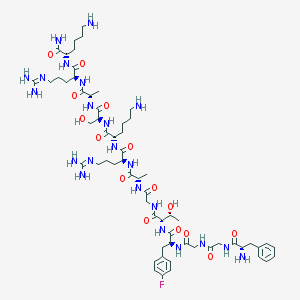 CHEMBL261997 CHEMBL261997 | C61H99FN22O15 | 1399.6 | 21 / 22 | -6.9 | No |
| 556413 | 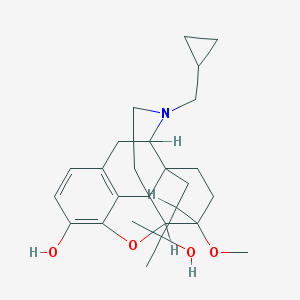 CHEMBL353979 CHEMBL353979 | C26H35NO4 | 425.569 | 5 / 2 | 3.6 | Yes |
| 243409 | 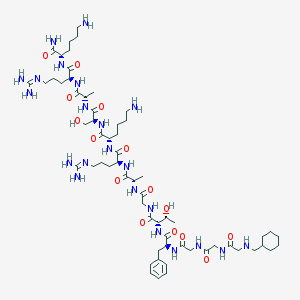 CHEMBL438234 CHEMBL438234 | C61H106N22O15 | 1387.66 | 20 / 22 | -6.1 | No |
| 243815 | 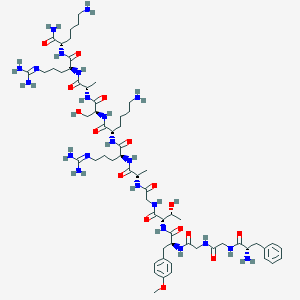 CHEMBL415583 CHEMBL415583 | C62H102N22O16 | 1411.64 | 21 / 22 | -7.0 | No |
| 245426 | 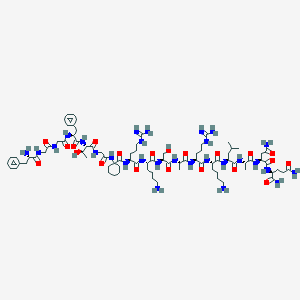 CHEMBL234724 CHEMBL234724 | C83H136N28O21 | 1862.18 | 26 / 30 | -7.4 | No |
| 245712 | 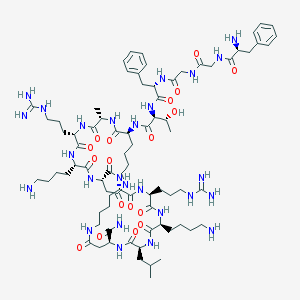 CID 49850693 CID 49850693 | C79H128N26O18 | 1730.06 | 23 / 27 | -4.7 | No |
| 246618 | 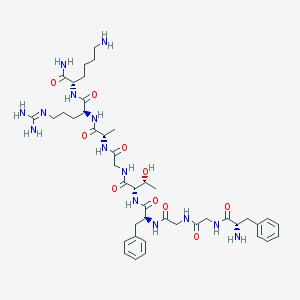 CHEMBL51556 CHEMBL51556 | C43H66N14O10 | 939.089 | 13 / 14 | -3.5 | No |
| 250209 | 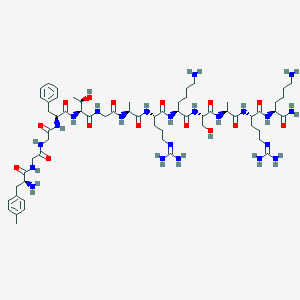 CHEMBL413178 CHEMBL413178 | C62H102N22O15 | 1395.64 | 20 / 22 | -6.6 | No |
| 556448 | 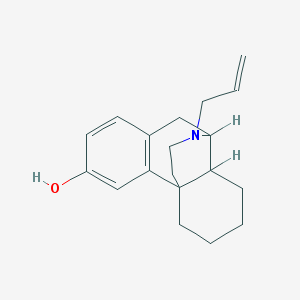 Morphinan-3-ol, 17-(2-propenyl)- Morphinan-3-ol, 17-(2-propenyl)- | C19H25NO | 283.415 | 2 / 1 | 3.5 | Yes |
| 556492 | 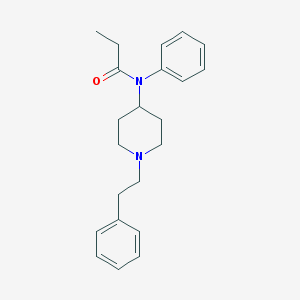 fentanyl fentanyl | C22H28N2O | 336.479 | 2 / 0 | 4.0 | Yes |
| 262956 | 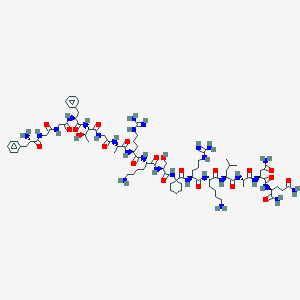 CHEMBL268394 CHEMBL268394 | C83H136N28O21 | 1862.18 | 26 / 30 | -6.8 | No |
| 264038 |  CID 53325329 CID 53325329 | C82H136FN29O21 | 1883.17 | 28 / 31 | -8.7 | No |
| 265616 |  CHEMBL398224 CHEMBL398224 | C82H136N28O21 | 1850.17 | 26 / 30 | -7.3 | No |
| 269179 |  BDBM21842 BDBM21842 | C79H129N27O22 | 1809.07 | 27 / 28 | -11.0 | No |
| 269322 |  D07WQG D07WQG | C79H128N26O23 | 1810.05 | 28 / 28 | -12.6 | No |
| 270190 |  Nociceptin(1-12)-NH2 Nociceptin(1-12)-NH2 | C55H88N20O14 | 1253.44 | 18 / 20 | -6.5 | No |
| 272630 |  CHEMBL415781 CHEMBL415781 | C82H134N28O21 | 1848.15 | 26 / 30 | -7.9 | No |
| 275181 |  CHEMBL233069 CHEMBL233069 | C80H132N28O21 | 1822.11 | 26 / 30 | -8.4 | No |
| 275505 |  CHEMBL437192 CHEMBL437192 | C84H140N28O21 | 1878.22 | 26 / 30 | -6.6 | No |
| 278272 | 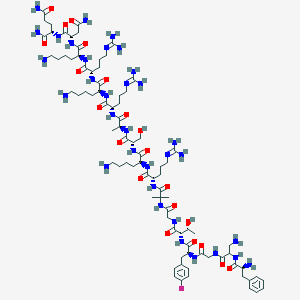 CHEMBL437744 CHEMBL437744 | C84H142FN33O21 | 1969.27 | 30 / 32 | -12.7 | No |
| 278278 | 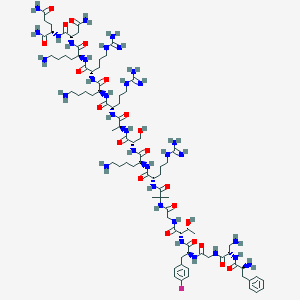 CHEMBL2372052 CHEMBL2372052 | C84H142FN33O21 | 1969.27 | 30 / 35 | -11.6 | No |
| 284898 | 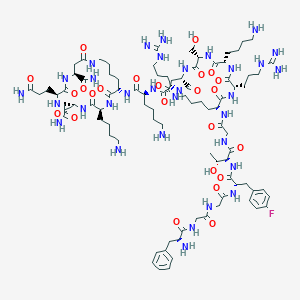 CID 49850695 CID 49850695 | C90H145FN32O24 | 2078.35 | 31 / 34 | -11.0 | No |
| 285675 | 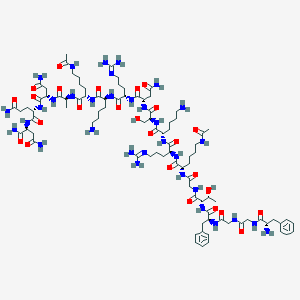 BDBM50333106 BDBM50333106 | C91H149N33O26 | 2121.4 | 31 / 33 | -13.9 | No |
| 295340 | 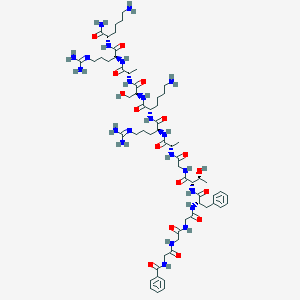 CHEMBL1631929 CHEMBL1631929 | C61H98N22O16 | 1395.59 | 20 / 22 | -7.0 | No |
| 296315 | 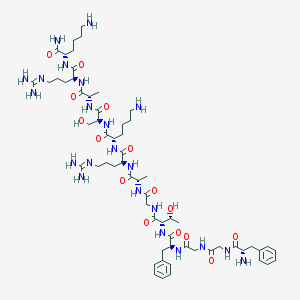 nociceptin(1-13)NH2 nociceptin(1-13)NH2 | C61H100N22O15 | 1381.61 | 20 / 22 | -7.0 | No |
| 299408 | 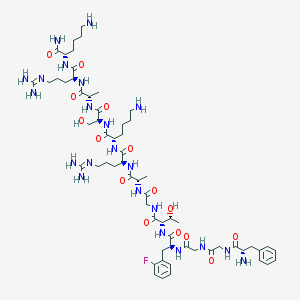 CHEMBL428668 CHEMBL428668 | C61H99FN22O15 | 1399.6 | 21 / 22 | -6.9 | No |
| 301313 | 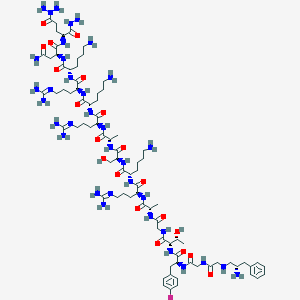 CHEMBL442297 CHEMBL442297 | C82H142FN35O20 | 1957.26 | 32 / 33 | -13.0 | No |
| 556678 | 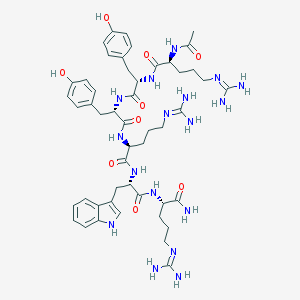 Ac-Arg-Tyr-Tyr-Arg-Trp-Arg-NH(2) Ac-Arg-Tyr-Tyr-Arg-Trp-Arg-NH(2) | C49H69N17O9 | 1040.2 | 12 / 16 | -2.3 | No |
| 307475 | 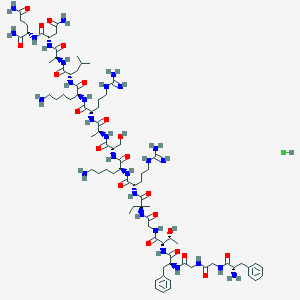 CHEMBL2372364 CHEMBL2372364 | C81H135ClN28O21 | 1872.6 | 26 / 31 | N/A | No |
| 312151 | 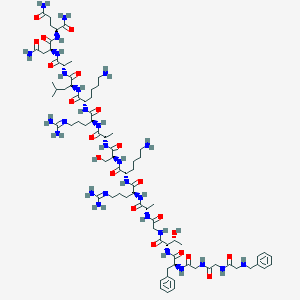 CHEMBL411621 CHEMBL411621 | C79H130N28O21 | 1808.08 | 26 / 28 | -9.3 | No |
| 323576 | 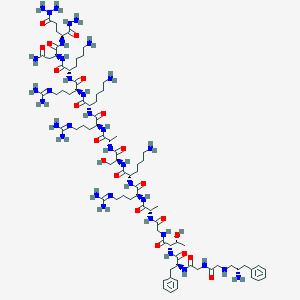 CHEMBL262544 CHEMBL262544 | C82H143N35O20 | 1939.27 | 31 / 33 | -13.1 | No |
| 556807 | 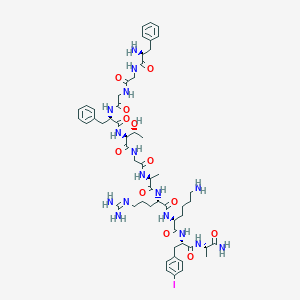 BDBM85823 BDBM85823 | C55H79IN16O12 | 1283.24 | 15 / 16 | -1.7 | No |
| 338135 | 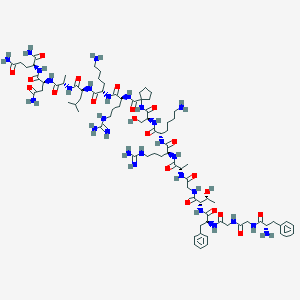 CHEMBL436819 CHEMBL436819 | C82H134N28O21 | 1848.15 | 26 / 30 | -7.4 | No |
| 340577 | 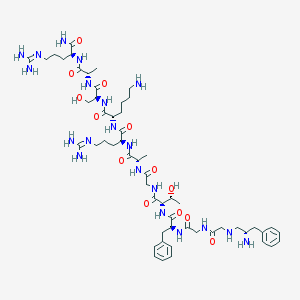 CHEMBL263859 CHEMBL263859 | C55H90N20O13 | 1239.45 | 18 / 20 | -6.2 | No |
| 341369 | 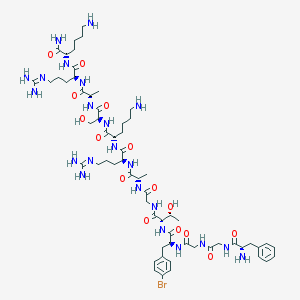 CHEMBL406097 CHEMBL406097 | C61H99BrN22O15 | 1460.51 | 20 / 22 | -6.3 | No |
| 556889 | 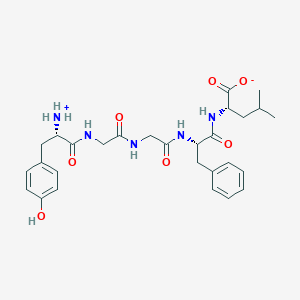 Leu-enkephalin Leu-enkephalin | C28H37N5O7 | 555.632 | 7 / 6 | -0.5 | No |
| 346578 | 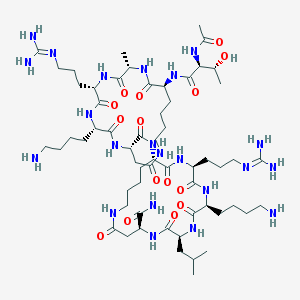 CHEMBL1631932 CHEMBL1631932 | C59H106N22O15 | 1363.64 | 19 / 21 | -6.5 | No |
| 347273 | 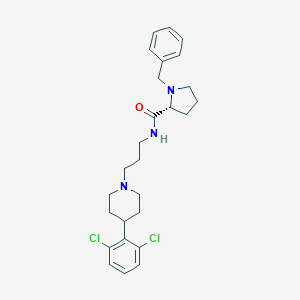 CHEMBL1783826 CHEMBL1783826 | C26H33Cl2N3O | 474.47 | 3 / 1 | 5.5 | No |
| 556908 | 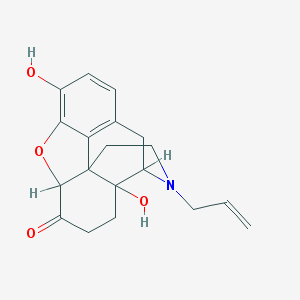 Naloxone(-) Naloxone(-) | C19H21NO4 | 327.38 | 5 / 2 | 2.1 | Yes |
| 351832 | 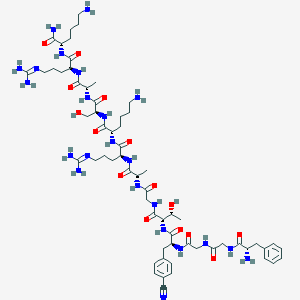 CHEMBL427791 CHEMBL427791 | C62H99N23O15 | 1406.62 | 21 / 22 | -7.3 | No |
| 354399 | 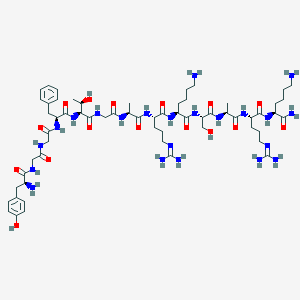 CHEMBL429363 CHEMBL429363 | C61H100N22O16 | 1397.61 | 21 / 23 | -7.3 | No |
| 356696 | 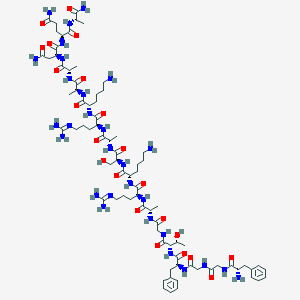 H-FGGFTGARKSARKAANQA-NH2 H-FGGFTGARKSARKAANQA-NH2 | C79H129N29O22 | 1837.08 | 27 / 29 | -10.9 | No |
| 556975 | 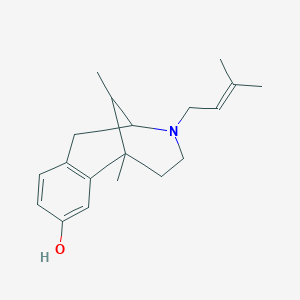 Fortalgesic Fortalgesic | C19H27NO | 285.431 | 2 / 1 | 3.3 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218