

You can:
| Name | Melanin-concentrating hormone receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MCHR1 |
| Synonym | SLC-1 Somatostatin receptor-like protein MCHR-1 MCHR MCH1R [ Show all ] |
| Disease | Obesity Obesity; Anxiety; Depression |
| Length | 422 |
| Amino acid sequence | MSVGAMKKGVGRAVGLGGGSGCQATEEDPLPNCGACAPGQGGRRWRLPQPAWVEGSSARLWEQATGTGWMDLEASLLPTGPNASNTSDGPDNLTSAGSPPRTGSISYINIIMPSVFGTICLLGIIGNSTVIFAVVKKSKLHWCNNVPDIFIINLSVVDLLFLLGMPFMIHQLMGNGVWHFGETMCTLITAMDANSQFTSTYILTAMAIDRYLATVHPISSTKFRKPSVATLVICLLWALSFISITPVWLYARLIPFPGGAVGCGIRLPNPDTDLYWFTLYQFFLAFALPFVVITAAYVRILQRMTSSVAPASQRSIRLRTKRVTRTAIAICLVFFVCWAPYYVLQLTQLSISRPTLTFVYLYNAAISLGYANSCLNPFVYIVLCETFRKRLVLSVKPAAQGQLRAVSNAQTADEERTESKGT |
| UniProt | Q99705 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q99705 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q99705. |
| BioLiP | N/A |
| Therapeutic Target Database | T09572 |
| ChEMBL | CHEMBL344 |
| IUPHAR | 280 |
| DrugBank | BE0003478 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 23 | 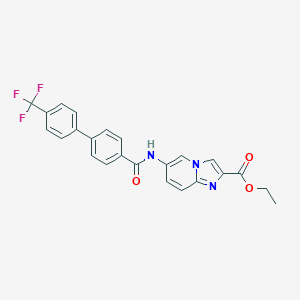 CHEMBL562537 CHEMBL562537 | C24H18F3N3O3 | 453.421 | 7 / 1 | 5.7 | No |
| 39 | 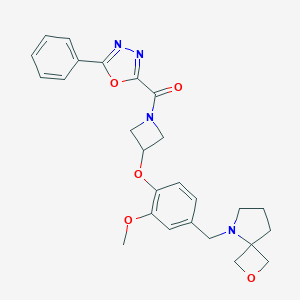 CHEMBL3670649 CHEMBL3670649 | C26H28N4O5 | 476.533 | 8 / 0 | 2.5 | Yes |
| 115 | 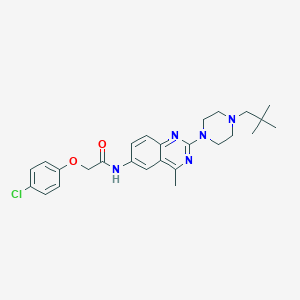 CHEMBL2016700 CHEMBL2016700 | C26H32ClN5O2 | 482.025 | 6 / 1 | 5.4 | No |
| 117 | 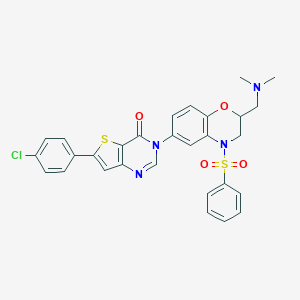 CHEMBL215374 CHEMBL215374 | C29H25ClN4O4S2 | 593.113 | 8 / 0 | 5.3 | No |
| 349 | 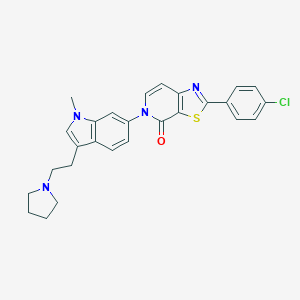 CHEMBL469343 CHEMBL469343 | C27H25ClN4OS | 489.034 | 4 / 0 | 5.6 | No |
| 412 | 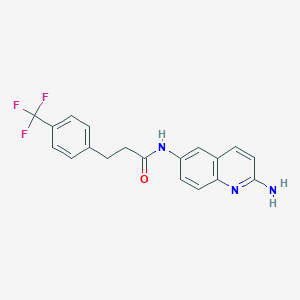 CHEMBL217751 CHEMBL217751 | C19H16F3N3O | 359.352 | 6 / 2 | 3.8 | Yes |
| 422 | 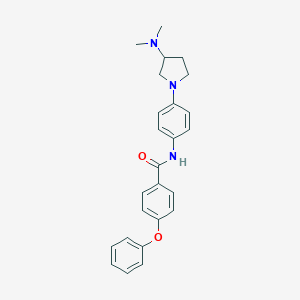 CHEMBL179870 CHEMBL179870 | C25H27N3O2 | 401.51 | 4 / 1 | 4.6 | Yes |
| 863 | 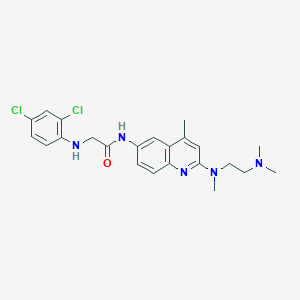 CHEMBL203804 CHEMBL203804 | C23H27Cl2N5O | 460.403 | 5 / 2 | 5.2 | No |
| 1377 | 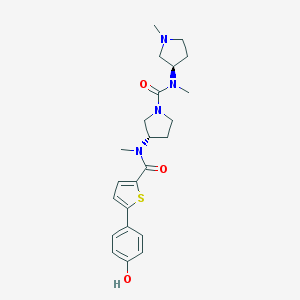 CHEMBL366275 CHEMBL366275 | C23H30N4O3S | 442.578 | 5 / 1 | 2.6 | Yes |
| 1403 | 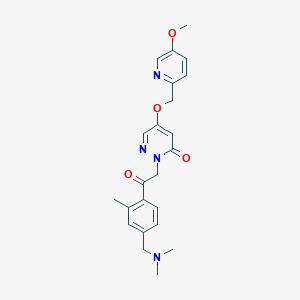 CHEMBL3601377 CHEMBL3601377 | C23H26N4O4 | 422.485 | 7 / 0 | 1.7 | Yes |
| 1570 | 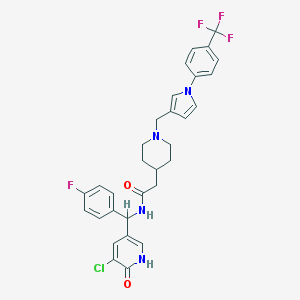 CHEMBL540930 CHEMBL540930 | C31H29ClF4N4O2 | 601.043 | 7 / 2 | 5.0 | No |
| 1691 | 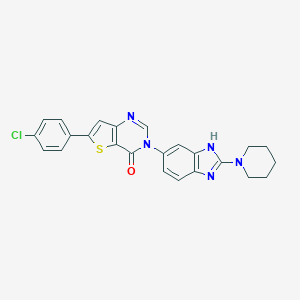 CHEMBL387217 CHEMBL387217 | C24H20ClN5OS | 461.968 | 5 / 1 | 5.6 | No |
| 1871 | 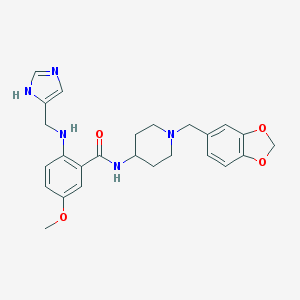 CHEMBL363169 CHEMBL363169 | C25H29N5O4 | 463.538 | 7 / 3 | 3.3 | Yes |
| 1879 | 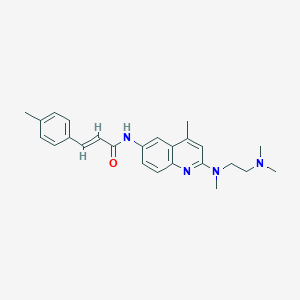 CHEMBL202563 CHEMBL202563 | C25H30N4O | 402.542 | 4 / 1 | 4.7 | Yes |
| 2237 | 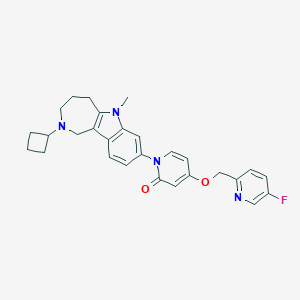 CHEMBL3665377 CHEMBL3665377 | C28H29FN4O2 | 472.564 | 5 / 0 | 3.4 | Yes |
| 2939 | 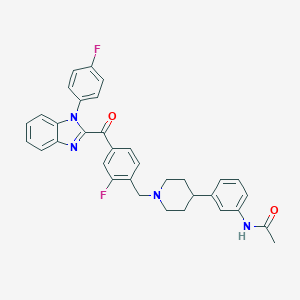 CHEMBL3219269 CHEMBL3219269 | C34H30F2N4O2 | 564.637 | 6 / 1 | 6.2 | No |
| 2965 | 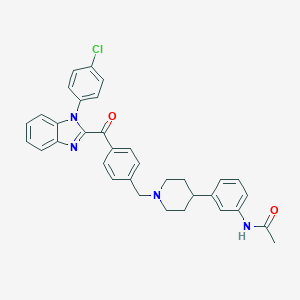 CHEMBL3219001 CHEMBL3219001 | C34H31ClN4O2 | 563.098 | 4 / 1 | 6.7 | No |
| 3083 | 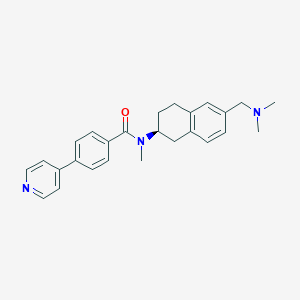 CHEMBL216050 CHEMBL216050 | C26H29N3O | 399.538 | 3 / 0 | 4.2 | Yes |
| 3195 | 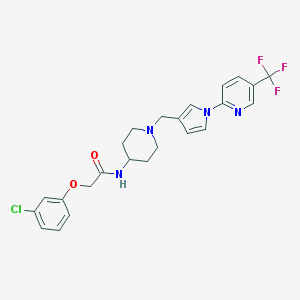 CHEMBL471514 CHEMBL471514 | C24H24ClF3N4O2 | 492.927 | 7 / 1 | 4.5 | Yes |
| 3638 | 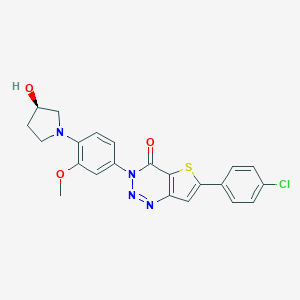 CHEMBL469790 CHEMBL469790 | C22H19ClN4O3S | 454.929 | 7 / 1 | 5.0 | Yes |
| 3662 | 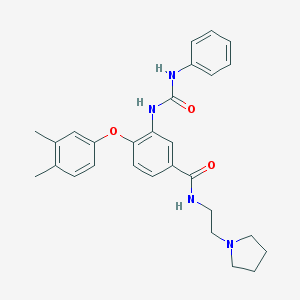 CHEMBL216192 CHEMBL216192 | C28H32N4O3 | 472.589 | 4 / 3 | 4.5 | Yes |
| 521546 | 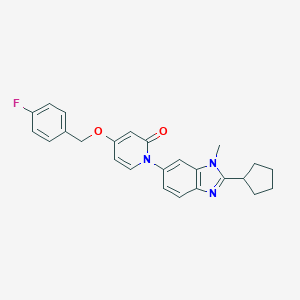 CHEMBL3793395 CHEMBL3793395 | C25H24FN3O2 | 417.484 | 4 / 0 | 4.4 | Yes |
| 3892 | 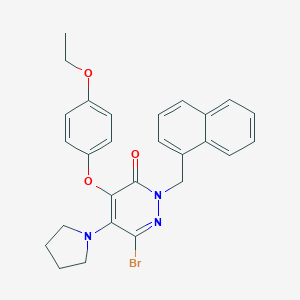 CHEMBL1995837 CHEMBL1995837 | C27H26BrN3O3 | 520.427 | 5 / 0 | 6.2 | No |
| 4030 | 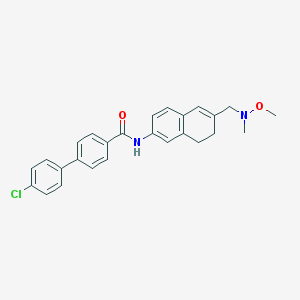 CHEMBL1818796 CHEMBL1818796 | C26H25ClN2O2 | 432.948 | 3 / 1 | 5.5 | No |
| 4109 | 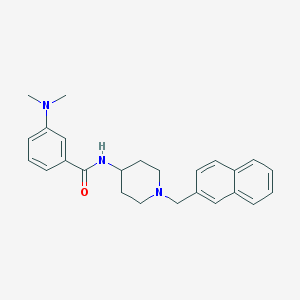 CHEMBL274102 CHEMBL274102 | C25H29N3O | 387.527 | 3 / 1 | 4.7 | Yes |
| 4400 | 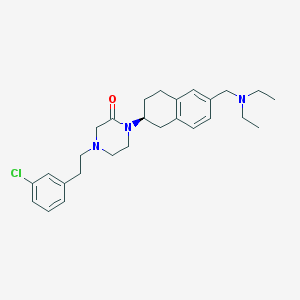 CHEMBL229792 CHEMBL229792 | C27H36ClN3O | 454.055 | 3 / 0 | 5.1 | No |
| 4719 | 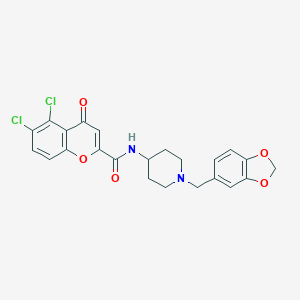 CHEMBL2113245 CHEMBL2113245 | C23H20Cl2N2O5 | 475.322 | 6 / 1 | 4.2 | Yes |
| 4814 | 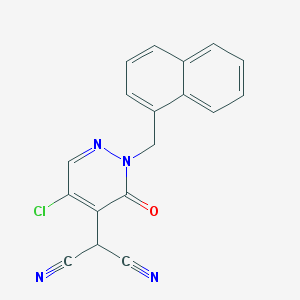 CHEMBL2001833 CHEMBL2001833 | C18H11ClN4O | 334.763 | 4 / 0 | 2.7 | Yes |
| 4819 | 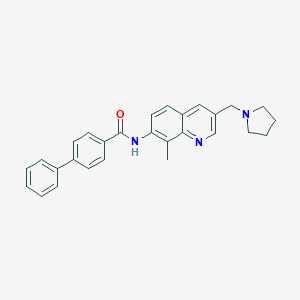 CHEMBL2031733 CHEMBL2031733 | C28H27N3O | 421.544 | 3 / 1 | 5.3 | No |
| 5009 | 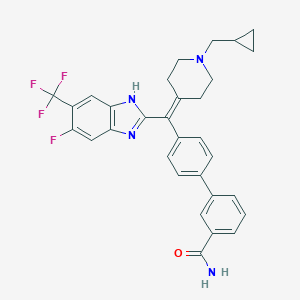 CHEMBL377270 CHEMBL377270 | C31H28F4N4O | 548.586 | 7 / 2 | 6.2 | No |
| 5120 | 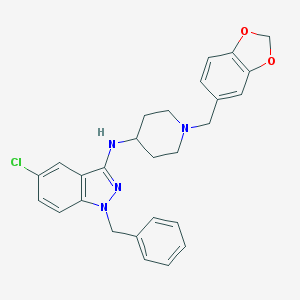 CHEMBL370576 CHEMBL370576 | C27H27ClN4O2 | 474.989 | 5 / 1 | 6.0 | No |
| 5167 | 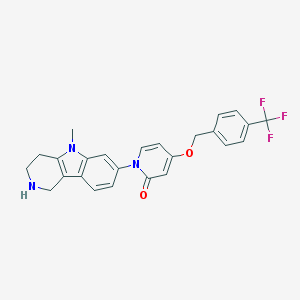 CHEMBL1290382 CHEMBL1290382 | C25H22F3N3O2 | 453.465 | 6 / 1 | 3.5 | Yes |
| 521611 | 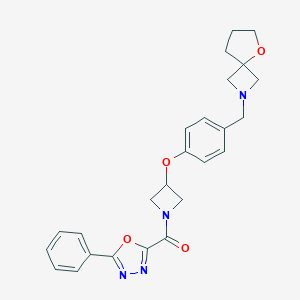 CHEMBL3786346 CHEMBL3786346 | C25H26N4O4 | 446.507 | 7 / 0 | 2.5 | Yes |
| 5357 | 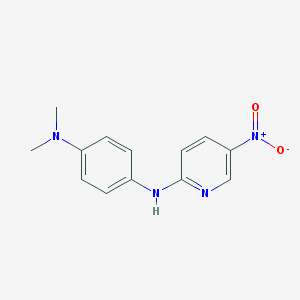 BAS 03293908 BAS 03293908 | C13H14N4O2 | 258.281 | 5 / 1 | 2.7 | Yes |
| 5388 | 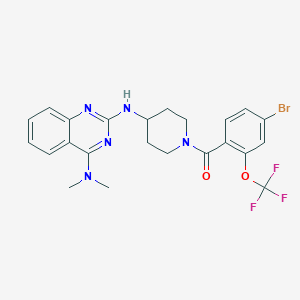 CHEMBL181251 CHEMBL181251 | C23H23BrF3N5O2 | 538.369 | 9 / 1 | 5.9 | No |
| 5444 | 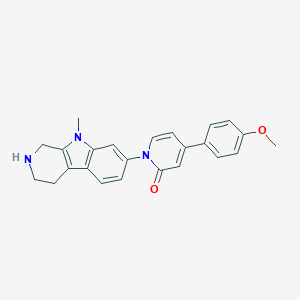 CHEMBL1289723 CHEMBL1289723 | C24H23N3O2 | 385.467 | 3 / 1 | 2.6 | Yes |
| 5458 | 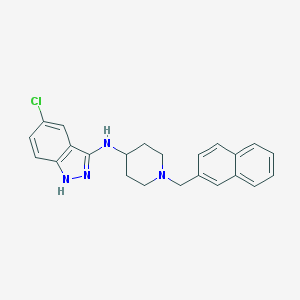 CHEMBL380782 CHEMBL380782 | C23H23ClN4 | 390.915 | 3 / 2 | 5.8 | No |
| 5554 | 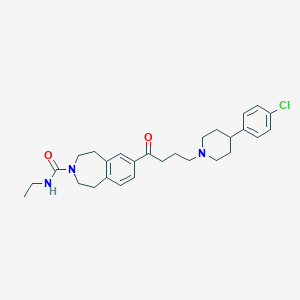 CHEMBL1914639 CHEMBL1914639 | C28H36ClN3O2 | 482.065 | 3 / 1 | 4.8 | Yes |
| 5832 | 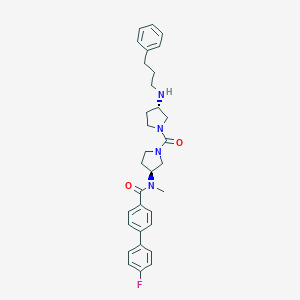 CHEMBL212715 CHEMBL212715 | C32H37FN4O2 | 528.672 | 4 / 1 | 4.9 | No |
| 463590 | 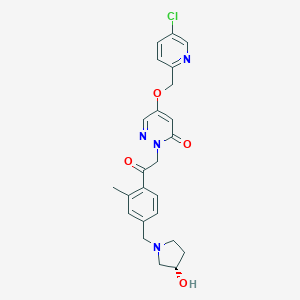 US8933079, 7.13 US8933079, 7.13 | C24H25ClN4O4 | 468.938 | 7 / 1 | 1.9 | Yes |
| 557442 | 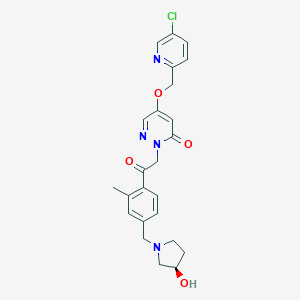 BDBM142208 BDBM142208 | C24H25ClN4O4 | 468.938 | 7 / 1 | 1.9 | Yes |
| 6111 | 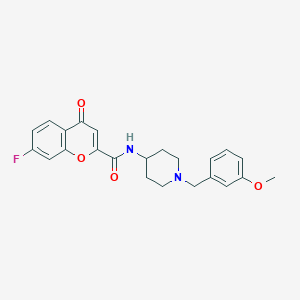 CHEMBL217400 CHEMBL217400 | C23H23FN2O4 | 410.445 | 6 / 1 | 3.2 | Yes |
| 6228 | 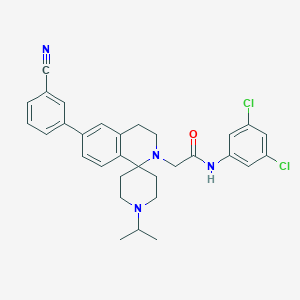 CHEMBL213491 CHEMBL213491 | C31H32Cl2N4O | 547.524 | 4 / 1 | 6.5 | No |
| 6232 | 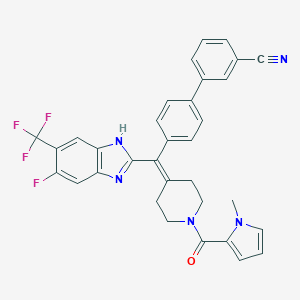 CHEMBL268938 CHEMBL268938 | C33H25F4N5O | 583.591 | 7 / 1 | 6.5 | No |
| 6352 | 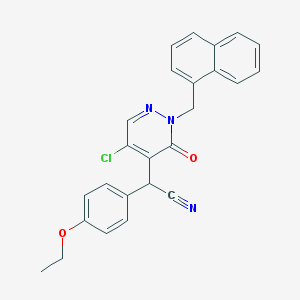 CHEMBL1973528 CHEMBL1973528 | C25H20ClN3O2 | 429.904 | 4 / 0 | 4.7 | Yes |
| 6394 | 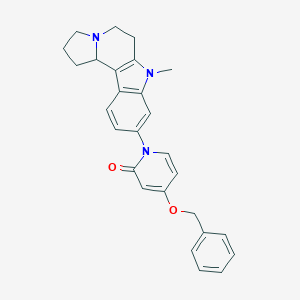 SCHEMBL94215 SCHEMBL94215 | C27H27N3O2 | 425.532 | 3 / 0 | 3.6 | Yes |
| 6460 | 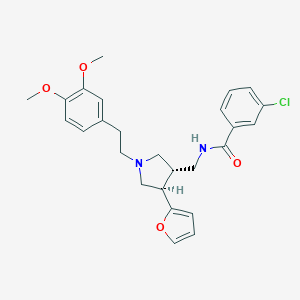 CHEMBL1760086 CHEMBL1760086 | C26H29ClN2O4 | 468.978 | 5 / 1 | 4.6 | Yes |
| 6471 | 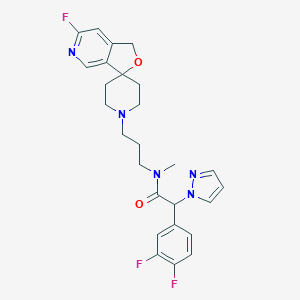 CHEMBL466425 CHEMBL466425 | C26H28F3N5O2 | 499.538 | 8 / 0 | 2.8 | Yes |
| 6517 | 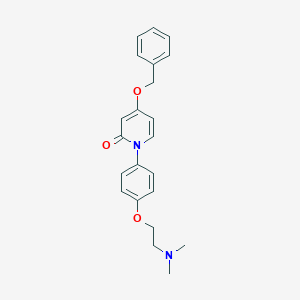 CHEMBL1642484 CHEMBL1642484 | C22H24N2O3 | 364.445 | 4 / 0 | 3.1 | Yes |
| 6545 | 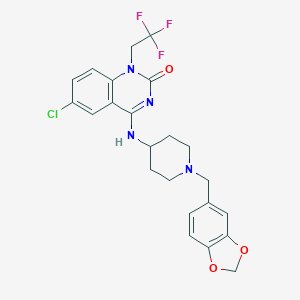 CHEMBL205492 CHEMBL205492 | C23H22ClF3N4O3 | 494.899 | 7 / 1 | 4.3 | Yes |
| 6728 | 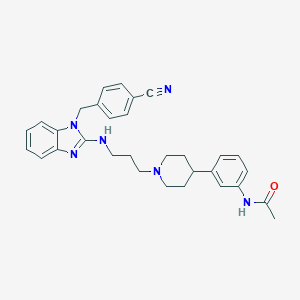 CHEMBL1223966 CHEMBL1223966 | C31H34N6O | 506.654 | 5 / 2 | 4.9 | No |
| 6753 | 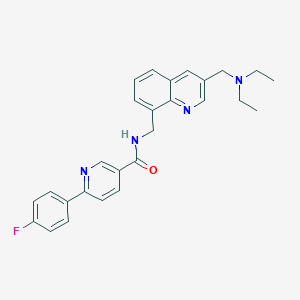 CHEMBL213817 CHEMBL213817 | C27H27FN4O | 442.538 | 5 / 1 | 4.2 | Yes |
| 6795 | 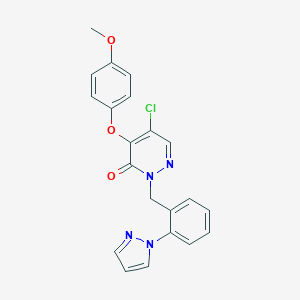 CHEMBL1981304 CHEMBL1981304 | C21H17ClN4O3 | 408.842 | 5 / 0 | 3.5 | Yes |
| 6956 | 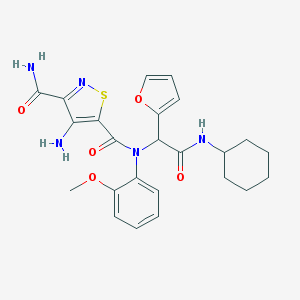 MLS001225295 MLS001225295 | C24H27N5O5S | 497.57 | 8 / 3 | 3.3 | Yes |
| 7038 | 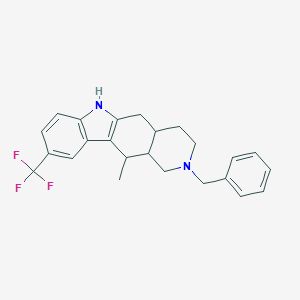 CHEMBL1922259 CHEMBL1922259 | C24H25F3N2 | 398.473 | 4 / 1 | 5.8 | No |
| 7149 | 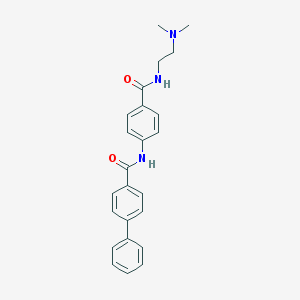 CHEMBL186601 CHEMBL186601 | C24H25N3O2 | 387.483 | 3 / 2 | 3.7 | Yes |
| 7342 | 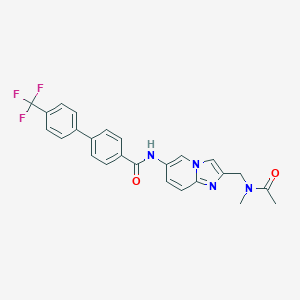 CHEMBL551408 CHEMBL551408 | C25H21F3N4O2 | 466.464 | 6 / 1 | 4.4 | Yes |
| 7370 | 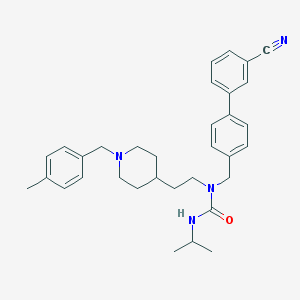 CHEMBL240116 CHEMBL240116 | C33H40N4O | 508.71 | 3 / 1 | 6.2 | No |
| 7492 | 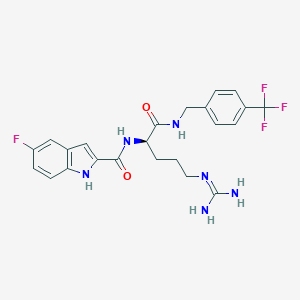 CHEMBL263171 CHEMBL263171 | C23H24F4N6O2 | 492.479 | 7 / 5 | 2.7 | Yes |
| 7516 | 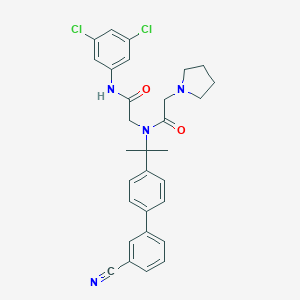 CHEMBL198724 CHEMBL198724 | C30H30Cl2N4O2 | 549.496 | 4 / 1 | 5.9 | No |
| 7592 | 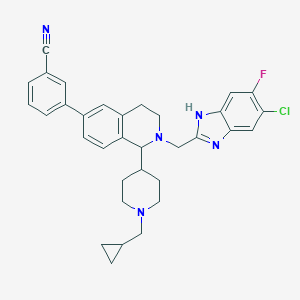 CHEMBL266840 CHEMBL266840 | C33H33ClFN5 | 554.11 | 5 / 1 | 6.5 | No |
| 7599 | 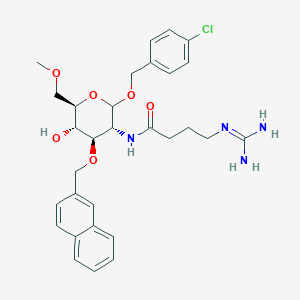 CHEMBL1254711 CHEMBL1254711 | C30H37ClN4O6 | 585.098 | 7 / 4 | 2.1 | No |
| 7600 | 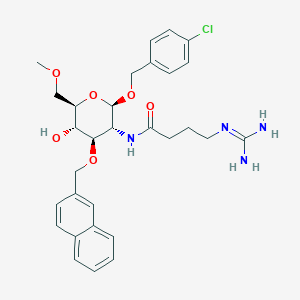 CHEMBL1254050 CHEMBL1254050 | C30H37ClN4O6 | 585.098 | 7 / 4 | 2.1 | No |
| 7601 | 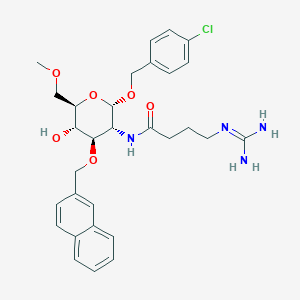 CHEMBL1253956 CHEMBL1253956 | C30H37ClN4O6 | 585.098 | 7 / 4 | 2.1 | No |
| 7713 | 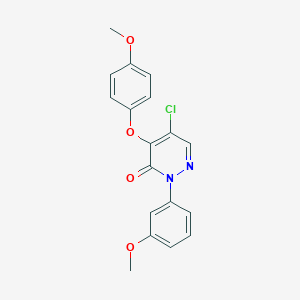 SR-02000000483 SR-02000000483 | C18H15ClN2O4 | 358.778 | 5 / 0 | 3.5 | Yes |
| 7857 | 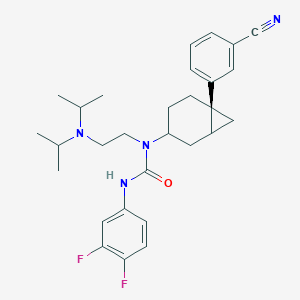 CHEMBL1254479 CHEMBL1254479 | C29H36F2N4O | 494.631 | 5 / 1 | 5.9 | No |
| 7858 | 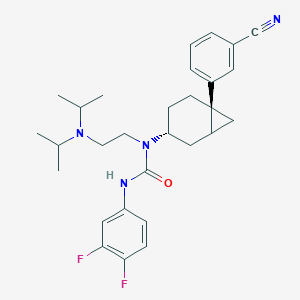 CHEMBL229769 CHEMBL229769 | C29H36F2N4O | 494.631 | 5 / 1 | 5.9 | No |
| 7873 | 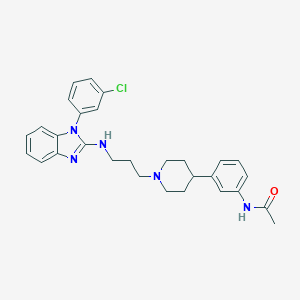 CHEMBL1223709 CHEMBL1223709 | C29H32ClN5O | 502.059 | 4 / 2 | 5.9 | No |
| 7887 | 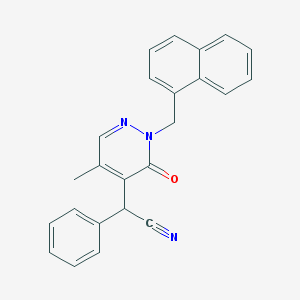 CHEMBL2003269 CHEMBL2003269 | C24H19N3O | 365.436 | 3 / 0 | 3.9 | Yes |
| 7935 | 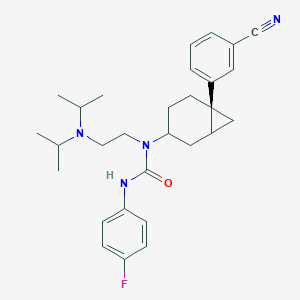 CHEMBL1254480 CHEMBL1254480 | C29H37FN4O | 476.64 | 4 / 1 | 5.8 | No |
| 7936 | 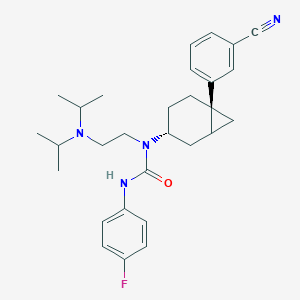 CHEMBL229822 CHEMBL229822 | C29H37FN4O | 476.64 | 4 / 1 | 5.8 | No |
| 8038 | 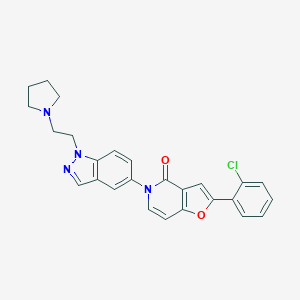 CHEMBL1290384 CHEMBL1290384 | C26H23ClN4O2 | 458.946 | 4 / 0 | 4.6 | Yes |
| 8252 | 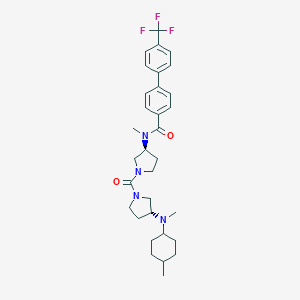 CHEMBL385959 CHEMBL385959 | C32H41F3N4O2 | 570.701 | 6 / 0 | 5.9 | No |
| 521705 | 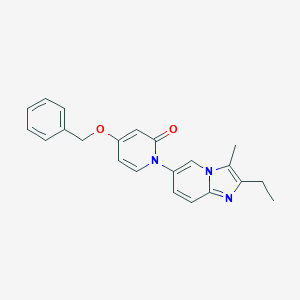 CHEMBL3771000 CHEMBL3771000 | C22H21N3O2 | 359.429 | 3 / 0 | 4.2 | Yes |
| 8752 | 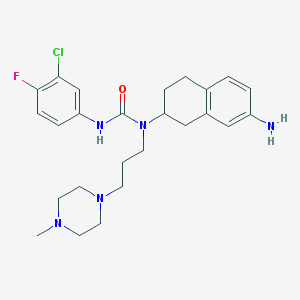 CHEMBL426339 CHEMBL426339 | C25H33ClFN5O | 474.021 | 5 / 2 | 3.8 | Yes |
| 8987 | 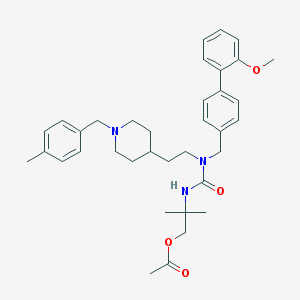 CHEMBL392013 CHEMBL392013 | C36H47N3O4 | 585.789 | 5 / 1 | 6.1 | No |
| 9150 | 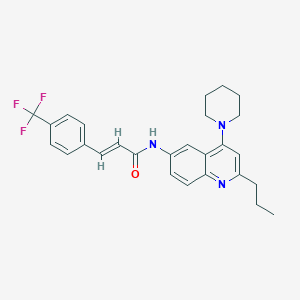 CHEMBL213873 CHEMBL213873 | C27H28F3N3O | 467.536 | 6 / 1 | 6.5 | No |
| 9174 | 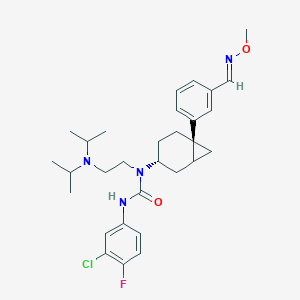 CHEMBL434747 CHEMBL434747 | C30H40ClFN4O2 | 543.124 | 5 / 1 | 6.8 | No |
| 9176 | 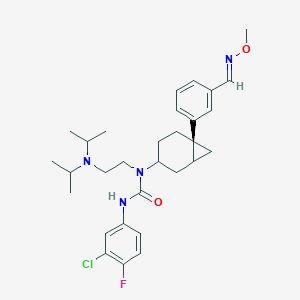 CHEMBL1254972 CHEMBL1254972 | C30H40ClFN4O2 | 543.124 | 5 / 1 | 6.8 | No |
| 9300 | 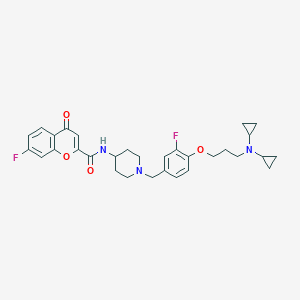 CHEMBL247891 CHEMBL247891 | C31H35F2N3O4 | 551.635 | 8 / 1 | 4.8 | No |
| 9306 | 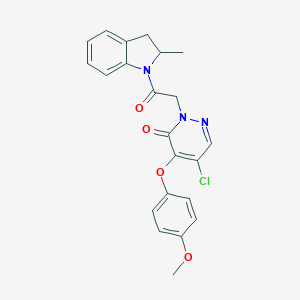 CHEMBL1968640 CHEMBL1968640 | C22H20ClN3O4 | 425.869 | 5 / 0 | 3.6 | Yes |
| 464011 | 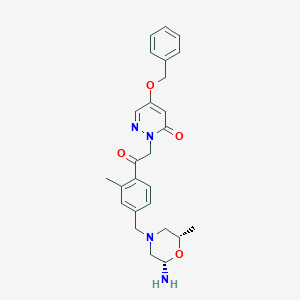 CHEMBL3686825 CHEMBL3686825 | C26H30N4O4 | 462.55 | 7 / 1 | 2.0 | Yes |
| 9437 | 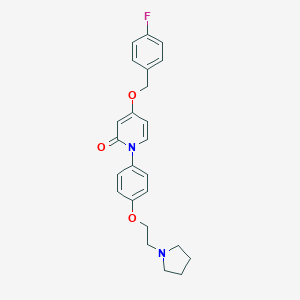 CHEMBL584538 CHEMBL584538 | C24H25FN2O3 | 408.473 | 5 / 0 | 3.7 | Yes |
| 9486 | 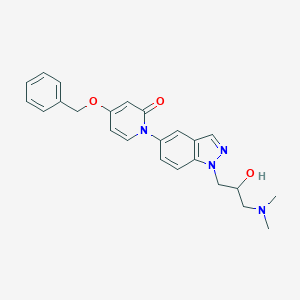 CHEMBL1289400 CHEMBL1289400 | C24H26N4O3 | 418.497 | 5 / 1 | 2.1 | Yes |
| 9518 | 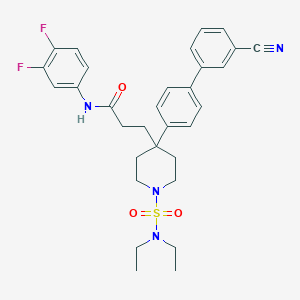 CHEMBL212671 CHEMBL212671 | C31H34F2N4O3S | 580.695 | 8 / 1 | 5.1 | No |
| 9523 | 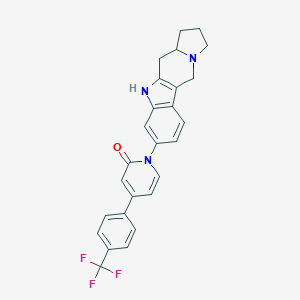 SCHEMBL173062 SCHEMBL173062 | C26H22F3N3O | 449.477 | 5 / 1 | 4.6 | Yes |
| 9636 | 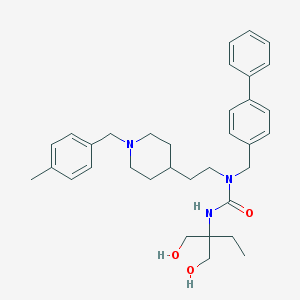 CHEMBL240287 CHEMBL240287 | C34H45N3O3 | 543.752 | 4 / 3 | 5.1 | No |
| 464059 | 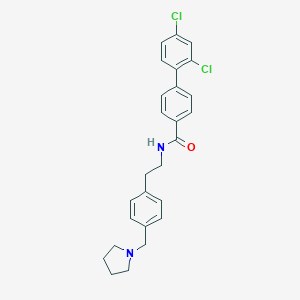 CHEMBL3600813 CHEMBL3600813 | C26H26Cl2N2O | 453.407 | 2 / 1 | 6.4 | No |
| 9716 | 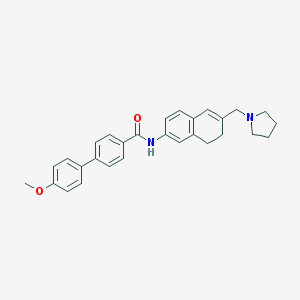 CHEMBL1818811 CHEMBL1818811 | C29H30N2O2 | 438.571 | 3 / 1 | 5.3 | No |
| 9775 | 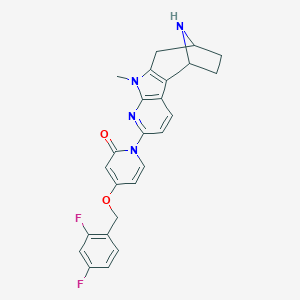 SCHEMBL155519 SCHEMBL155519 | C25H22F2N4O2 | 448.474 | 6 / 1 | 3.0 | Yes |
| 9832 | 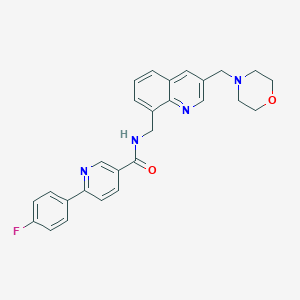 CHEMBL215131 CHEMBL215131 | C27H25FN4O2 | 456.521 | 6 / 1 | 3.1 | Yes |
| 10229 | 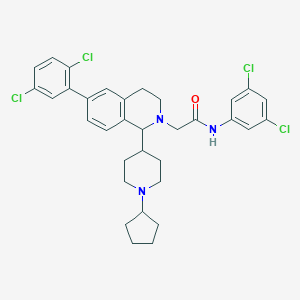 CHEMBL384116 CHEMBL384116 | C33H35Cl4N3O | 631.463 | 3 / 1 | 8.9 | No |
| 10422 | 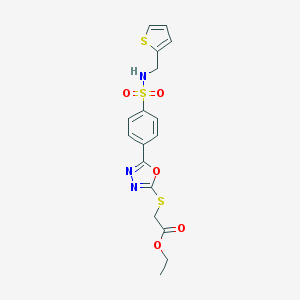 MLS000123364 MLS000123364 | C17H17N3O5S3 | 439.519 | 10 / 1 | 2.7 | Yes |
| 10615 | 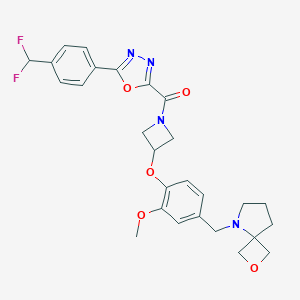 CHEMBL3670650 CHEMBL3670650 | C27H28F2N4O5 | 526.541 | 10 / 0 | 3.1 | No |
| 10635 | 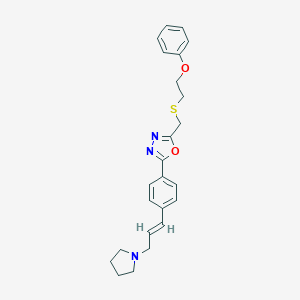 CHEMBL472485 CHEMBL472485 | C24H27N3O2S | 421.559 | 6 / 0 | 4.5 | Yes |
| 10871 | 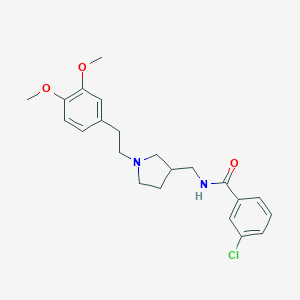 CHEMBL1760221 CHEMBL1760221 | C22H27ClN2O3 | 402.919 | 4 / 1 | 4.2 | Yes |
| 11037 | 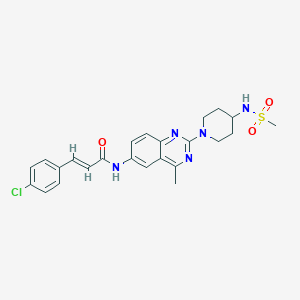 CHEMBL2017757 CHEMBL2017757 | C24H26ClN5O3S | 500.014 | 7 / 2 | 3.9 | No |
| 11286 | 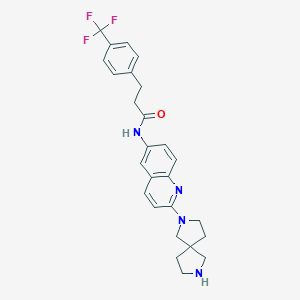 CHEMBL217502 CHEMBL217502 | C26H27F3N4O | 468.524 | 7 / 2 | 4.5 | Yes |
| 11311 | 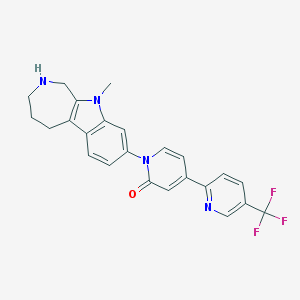 CHEMBL3665327 CHEMBL3665327 | C24H21F3N4O | 438.454 | 6 / 1 | 2.8 | Yes |
| 11594 | 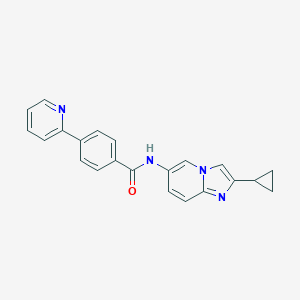 CHEMBL557317 CHEMBL557317 | C22H18N4O | 354.413 | 3 / 1 | 3.8 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218