

You can:
| Name | Metabotropic glutamate receptor 7 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | GRM7 |
| Synonym | GLUR7 glutamate receptor GPRC1G mGlu7 receptor mGlu7a receptor [ Show all ] |
| Disease | N/A |
| Length | 915 |
| Amino acid sequence | MVQLRKLLRVLTLMKFPCCVLEVLLCALAAAARGQEMYAPHSIRIEGDVTLGGLFPVHAKGPSGVPCGDIKRENGIHRLEAMLYALDQINSDPNLLPNVTLGARILDTCSRDTYALEQSLTFVQALIQKDTSDVRCTNGEPPVFVKPEKVVGVIGASGSSVSIMVANILRLFQIPQISYASTAPELSDDRRYDFFSRVVPPDSFQAQAMVDIVKALGWNYVSTLASEGSYGEKGVESFTQISKEAGGLCIAQSVRIPQERKDRTIDFDRIIKQLLDTPNSRAVVIFANDEDIKQILAAAKRADQVGHFLWVGSDSWGSKINPLHQHEDIAEGAITIQPKRATVEGFDAYFTSRTLENNRRNVWFAEYWEENFNCKLTISGSKKEDTDRKCTGQERIGKDSNYEQEGKVQFVIDAVYAMAHALHHMNKDLCADYRGVCPEMEQAGGKKLLKYIRNVNFNGSAGTPVMFNKNGDAPGRYDIFQYQTTNTSNPGYRLIGQWTDELQLNIEDMQWGKGVREIPASVCTLPCKPGQRKKTQKGTPCCWTCEPCDGYQYQFDEMTCQHCPYDQRPNENRTGCQDIPIIKLEWHSPWAVIPVFLAMLGIIATIFVMATFIRYNDTPIVRASGRELSYVLLTGIFLCYIITFLMIAKPDVAVCSFRRVFLGLGMCISYAALLTKTNRIYRIFEQGKKSVTAPRLISPTSQLAITSSLISVQLLGVFIWFGVDPPNIIIDYDEHKTMNPEQARGVLKCDITDLQIICSLGYSILLMVTCTVYAIKTRGVPENFNEAKPIGFTMYTTCIVWLAFIPIFFGTAQSAEKLYIQTTTLTISMNLSASVALGMLYMPKVYIIIFHPELNVQKRKRSFKAVVTAATMSSRLSHKPSDRPNGEAKTELCENVDPNSPAAKKKYVSYNNLVI |
| UniProt | Q14831 |
| Protein Data Bank | 5c5c, 3mq4 |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 5c5c. |
| BioLiP | BL0319784,BL0319785,BL0319786,, BL0181059, BL0181060, BL0319782,BL0319783 |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3777 |
| IUPHAR | 295 |
| DrugBank | BE0000834 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 8270 | 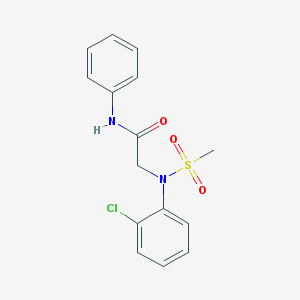 CHEMBL2208408 CHEMBL2208408 | C15H15ClN2O3S | 338.806 | 4 / 1 | 2.6 | Yes |
| 10403 | 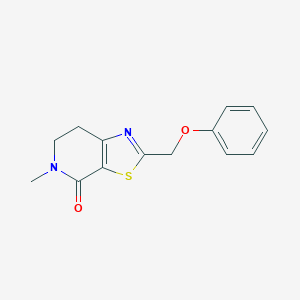 CHEMBL2426616 CHEMBL2426616 | C14H14N2O2S | 274.338 | 4 / 0 | 2.3 | Yes |
| 521769 | 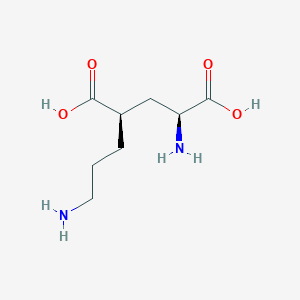 CHEMBL2312685 CHEMBL2312685 | C8H16N2O4 | 204.226 | 6 / 4 | -6.8 | Yes |
| 13325 | 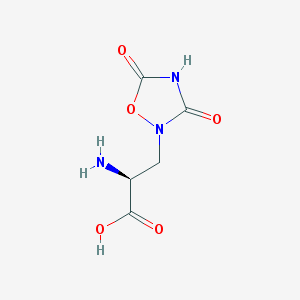 QUISQUALIC ACID QUISQUALIC ACID | C5H7N3O5 | 189.127 | 6 / 3 | -3.9 | Yes |
| 20315 | 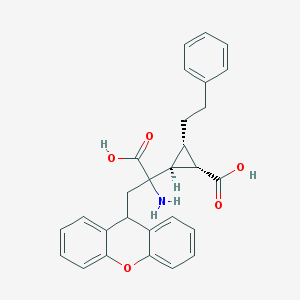 CHEMBL319279 CHEMBL319279 | C28H27NO5 | 457.526 | 6 / 3 | 2.1 | Yes |
| 465375 | 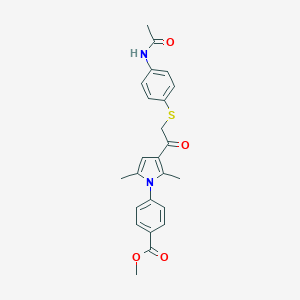 AC1N2AWG AC1N2AWG | C24H24N2O4S | 436.526 | 5 / 1 | 4.1 | Yes |
| 553357 | 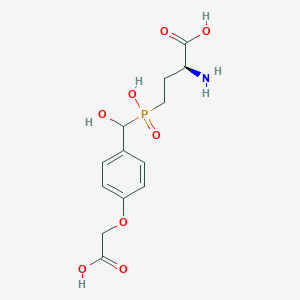 LSP4-2022 LSP4-2022 | C13H18NO8P | 347.26 | 9 / 5 | -3.8 | Yes |
| 466689 | 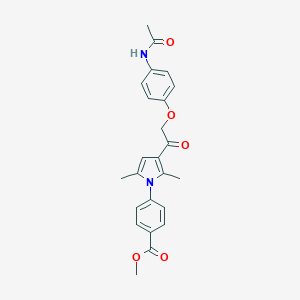 CHEMBL3560146 CHEMBL3560146 | C24H24N2O5 | 420.465 | 5 / 1 | 3.5 | Yes |
| 35790 | 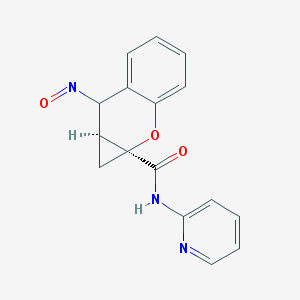 BDBM50388941 BDBM50388941 | C16H13N3O3 | 295.298 | 5 / 1 | 1.4 | Yes |
| 36683 | 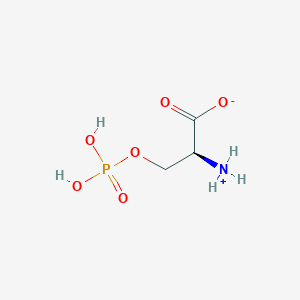 CID 57689797 CID 57689797 | C3H8NO6P | 185.072 | 6 / 3 | -4.5 | Yes |
| 36692 | 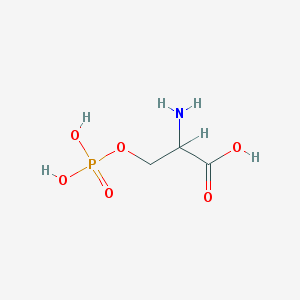 dl-O-Phosphoserine dl-O-Phosphoserine | C3H8NO6P | 185.072 | 7 / 4 | -5.1 | Yes |
| 467492 | 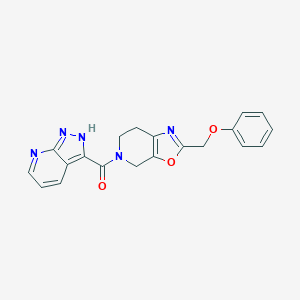 CHEMBL3605296 CHEMBL3605296 | C20H17N5O3 | 375.388 | 6 / 1 | 2.1 | Yes |
| 522873 | 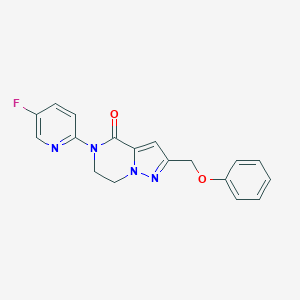 CHEMBL3746457 CHEMBL3746457 | C18H15FN4O2 | 338.342 | 5 / 0 | 2.2 | Yes |
| 553468 | 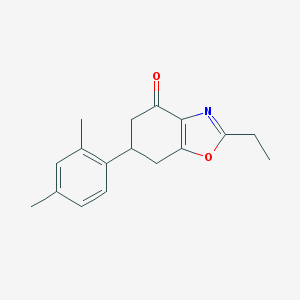 ADX-71743 ADX-71743 | C17H19NO2 | 269.344 | 3 / 0 | 3.8 | Yes |
| 469263 | 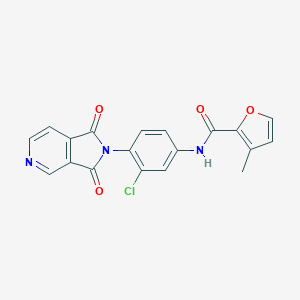 CHEMBL3628115 CHEMBL3628115 | C19H12ClN3O4 | 381.772 | 5 / 1 | 2.6 | Yes |
| 53132 | 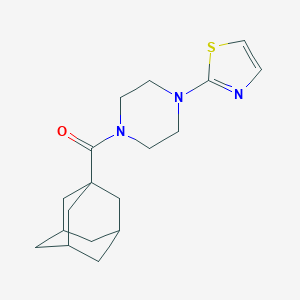 CHEMBL1688377 CHEMBL1688377 | C18H25N3OS | 331.478 | 4 / 0 | 3.3 | Yes |
| 55709 | 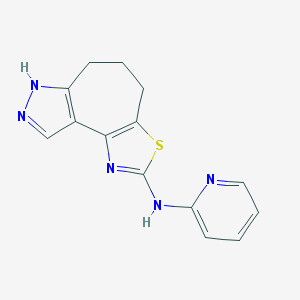 CHEMBL1830707 CHEMBL1830707 | C14H13N5S | 283.353 | 5 / 2 | 2.8 | Yes |
| 469952 | 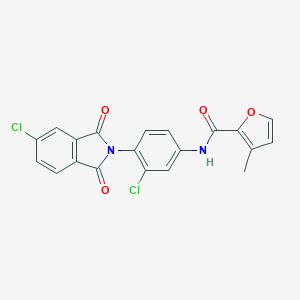 CHEMBL3628112 CHEMBL3628112 | C20H12Cl2N2O4 | 415.226 | 4 / 1 | 4.3 | Yes |
| 57276 | 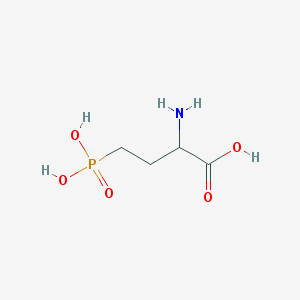 2-amino-4-phosphonobutanoic acid 2-amino-4-phosphonobutanoic acid | C4H10NO5P | 183.1 | 6 / 4 | -5.5 | Yes |
| 57290 | 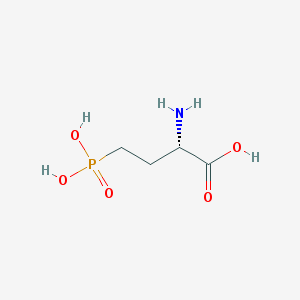 L-AP4 L-AP4 | C4H10NO5P | 183.1 | 6 / 4 | -5.5 | Yes |
| 61158 | 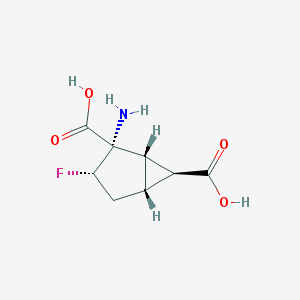 CHEMBL334014 CHEMBL334014 | C8H10FNO4 | 203.169 | 6 / 3 | -3.2 | Yes |
| 62856 | 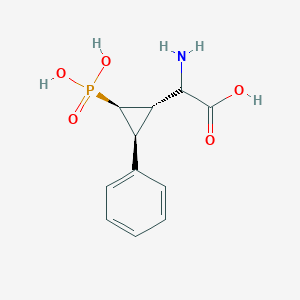 CHEMBL392419 CHEMBL392419 | C11H14NO5P | 271.209 | 6 / 4 | -3.3 | Yes |
| 62868 | 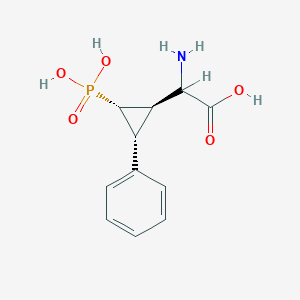 CHEMBL392420 CHEMBL392420 | C11H14NO5P | 271.209 | 6 / 4 | -3.3 | Yes |
| 553565 | 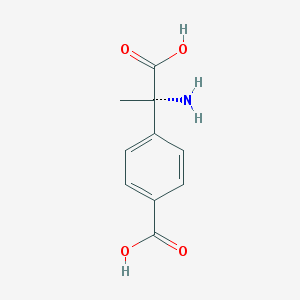 (S)-MCPG (S)-MCPG | C10H11NO4 | 209.201 | 5 / 3 | -2.0 | Yes |
| 471016 | 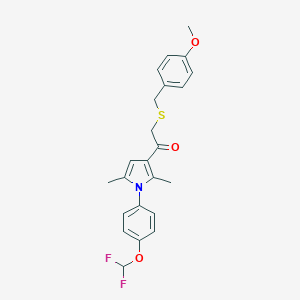 CHEMBL3561387 CHEMBL3561387 | C23H23F2NO3S | 431.498 | 6 / 0 | 5.9 | No |
| 553600 | 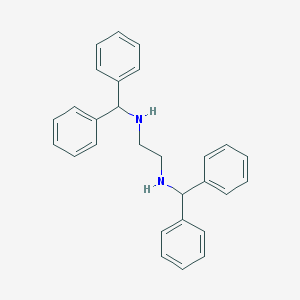 MLS001173561 MLS001173561 | C28H28N2 | 392.546 | 2 / 2 | 5.7 | No |
| 70681 | 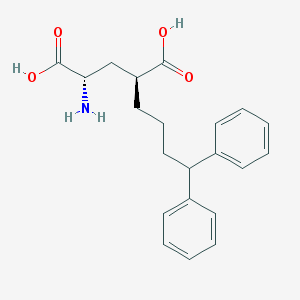 LY307452 LY307452 | C21H25NO4 | 355.434 | 5 / 3 | 1.4 | Yes |
| 79206 | 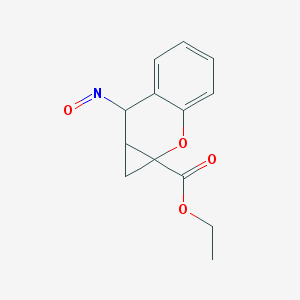 BDBM86213 BDBM86213 | C13H13NO4 | 247.25 | 5 / 0 | 1.5 | Yes |
| 80651 | 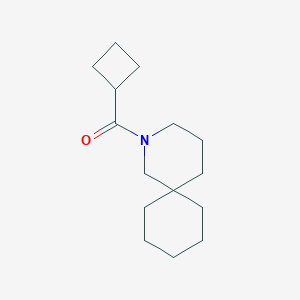 CHEMBL1783985 CHEMBL1783985 | C15H25NO | 235.371 | 1 / 0 | 3.4 | Yes |
| 473325 | 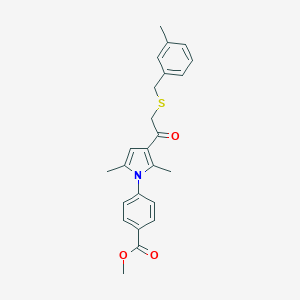 CHEMBL3561819 CHEMBL3561819 | C24H25NO3S | 407.528 | 4 / 0 | 5.2 | No |
| 89471 | 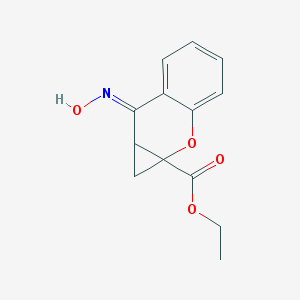 CPCCOEt CPCCOEt | C13H13NO4 | 247.25 | 5 / 1 | 1.9 | Yes |
| 553715 | 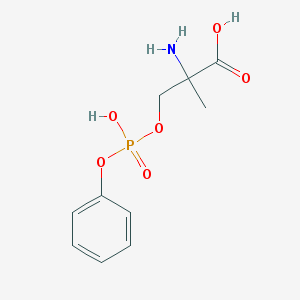 MSOPPE MSOPPE | C10H14NO6P | 275.197 | 7 / 3 | -2.6 | Yes |
| 524196 | 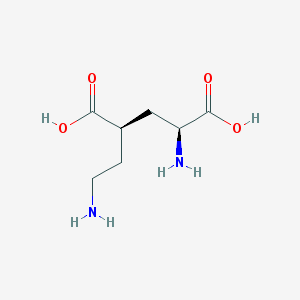 CHEMBL2312684 CHEMBL2312684 | C7H14N2O4 | 190.199 | 6 / 4 | -6.3 | Yes |
| 474766 | 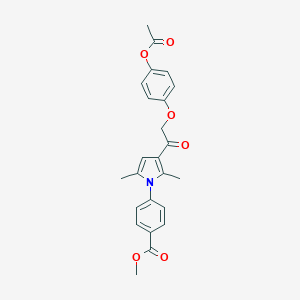 CHEMBL3560922 CHEMBL3560922 | C24H23NO6 | 421.449 | 6 / 0 | 4.1 | Yes |
| 96470 | 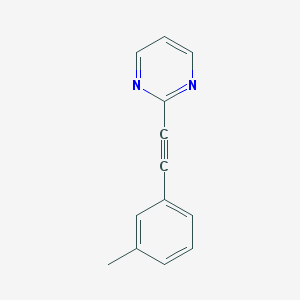 2-(m-tolylethynyl)pyrimidine 2-(m-tolylethynyl)pyrimidine | C13H10N2 | 194.237 | 2 / 0 | 2.6 | Yes |
| 99223 | 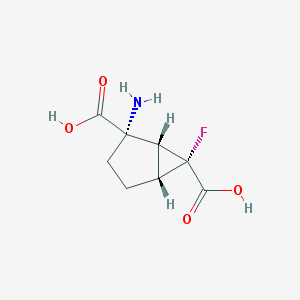 CHEMBL333519 CHEMBL333519 | C8H10FNO4 | 203.169 | 6 / 3 | -3.0 | Yes |
| 553765 | 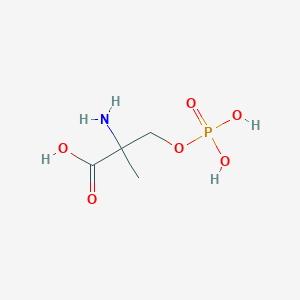 MSOP MSOP | C4H10NO6P | 199.099 | 7 / 4 | -5.0 | Yes |
| 553778 | 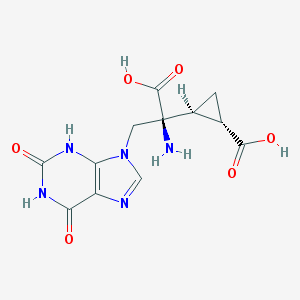 D0N9UZ D0N9UZ | C12H13N5O6 | 323.265 | 8 / 5 | -4.7 | Yes |
| 109295 | 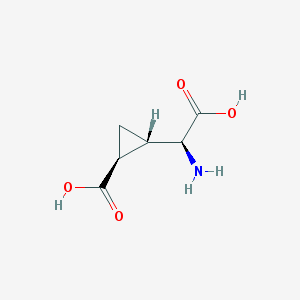 (2S,1'S,2'S)-2-(carboxycyclopropyl)glycine (2S,1'S,2'S)-2-(carboxycyclopropyl)glycine | C6H9NO4 | 159.141 | 5 / 3 | -3.4 | Yes |
| 477085 | 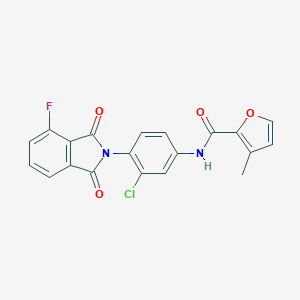 CHEMBL3628113 CHEMBL3628113 | C20H12ClFN2O4 | 398.774 | 5 / 1 | 3.8 | Yes |
| 524876 | 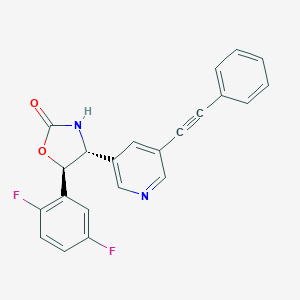 CHEMBL3804846 CHEMBL3804846 | C22H14F2N2O2 | 376.363 | 5 / 1 | 3.9 | Yes |
| 446356 | 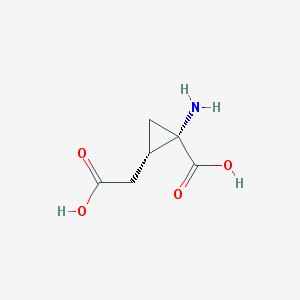 CTK0H1302 CTK0H1302 | C6H9NO4 | 159.141 | 5 / 3 | -3.7 | Yes |
| 446365 | 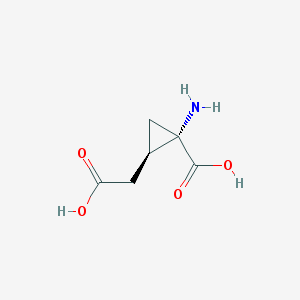 CHEMBL3347670 CHEMBL3347670 | C6H9NO4 | 159.141 | 5 / 3 | -3.7 | Yes |
| 119810 | 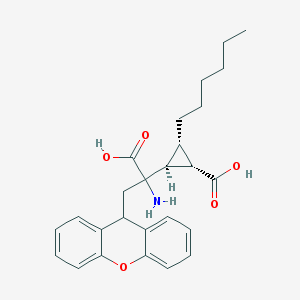 CHEMBL319732 CHEMBL319732 | C26H31NO5 | 437.536 | 6 / 3 | 2.9 | Yes |
| 553880 | 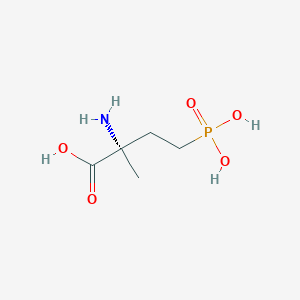 MAP4 MAP4 | C5H12NO5P | 197.127 | 6 / 4 | -4.6 | Yes |
| 120225 | 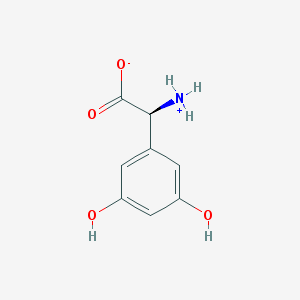 CHEBI:75204 CHEBI:75204 | C8H9NO4 | 183.163 | 4 / 3 | -1.8 | Yes |
| 127110 | 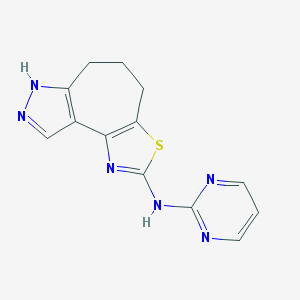 CHEMBL1830711 CHEMBL1830711 | C13H12N6S | 284.341 | 6 / 2 | 2.1 | Yes |
| 480598 | 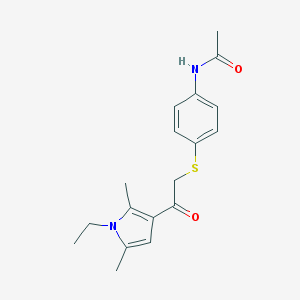 AC1P03JT AC1P03JT | C18H22N2O2S | 330.446 | 3 / 1 | 2.9 | Yes |
| 151148 | 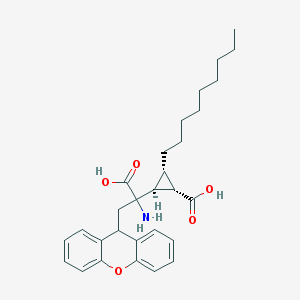 CHEMBL420262 CHEMBL420262 | C29H37NO5 | 479.617 | 6 / 3 | 4.5 | Yes |
| 482196 | 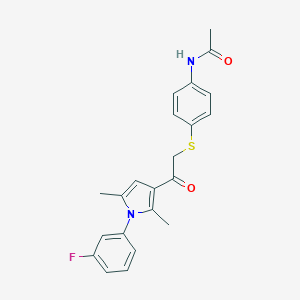 CHEMBL3561545 CHEMBL3561545 | C22H21FN2O2S | 396.48 | 4 / 1 | 4.3 | Yes |
| 158299 | 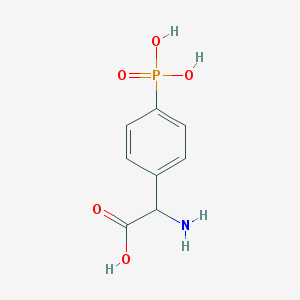 (RS)-PPG (RS)-PPG | C8H10NO5P | 231.144 | 6 / 4 | -3.7 | Yes |
| 482582 | 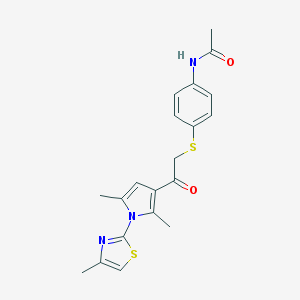 CHEMBL3561788 CHEMBL3561788 | C20H21N3O2S2 | 399.527 | 5 / 1 | 4.0 | Yes |
| 554125 | 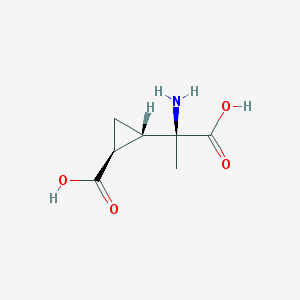 MCCG-I MCCG-I | C7H11NO4 | 173.168 | 5 / 3 | -3.3 | Yes |
| 167553 | 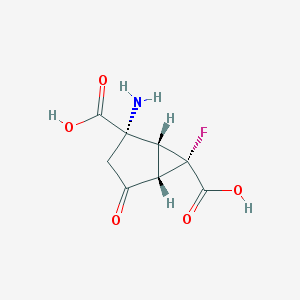 MGS-0028 MGS-0028 | C8H8FNO5 | 217.152 | 7 / 3 | -4.2 | Yes |
| 181959 | 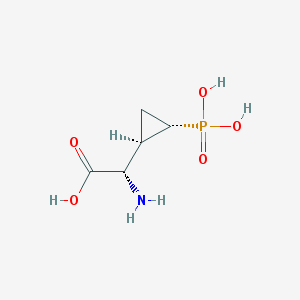 CHEMBL197976 CHEMBL197976 | C5H10NO5P | 195.111 | 6 / 4 | -4.7 | Yes |
| 181965 | 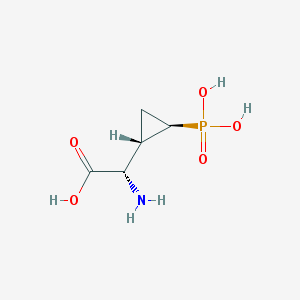 CHEMBL199626 CHEMBL199626 | C5H10NO5P | 195.111 | 6 / 4 | -4.7 | Yes |
| 181974 | 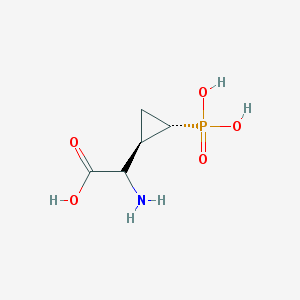 CHEMBL442076 CHEMBL442076 | C5H10NO5P | 195.111 | 6 / 4 | -4.7 | Yes |
| 526734 | 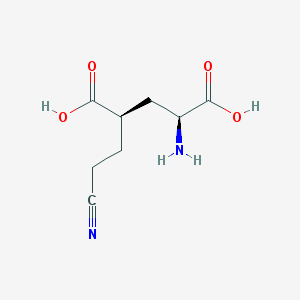 CHEMBL2312682 CHEMBL2312682 | C8H12N2O4 | 200.194 | 6 / 3 | -3.4 | Yes |
| 486353 | 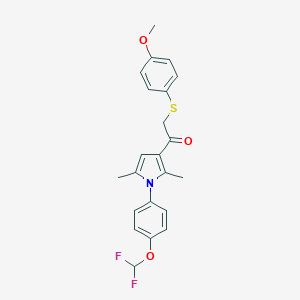 CHEMBL3561539 CHEMBL3561539 | C22H21F2NO3S | 417.471 | 6 / 0 | 5.9 | No |
| 563362 | 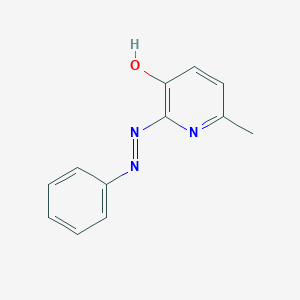 SIB 1757 SIB 1757 | C12H11N3O | 213.24 | 4 / 1 | 3.0 | Yes |
| 192706 | 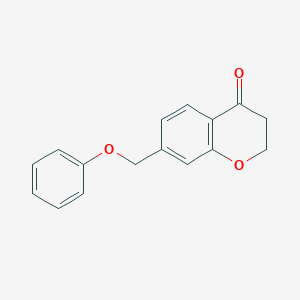 CHEMBL1682820 CHEMBL1682820 | C16H14O3 | 254.285 | 3 / 0 | 2.8 | Yes |
| 195662 | 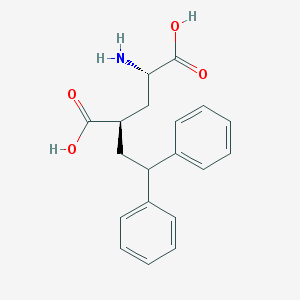 CHEMBL40123 CHEMBL40123 | C19H21NO4 | 327.38 | 5 / 3 | 0.3 | Yes |
| 201023 | 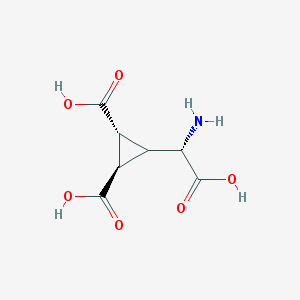 Dcg-IV Dcg-IV | C7H9NO6 | 203.15 | 7 / 4 | -4.3 | Yes |
| 488356 | 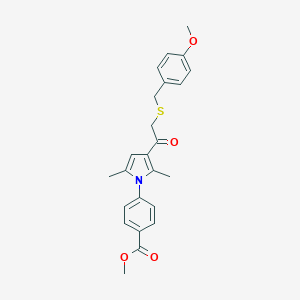 CHEMBL3560239 CHEMBL3560239 | C24H25NO4S | 423.527 | 5 / 0 | 4.8 | Yes |
| 488362 | 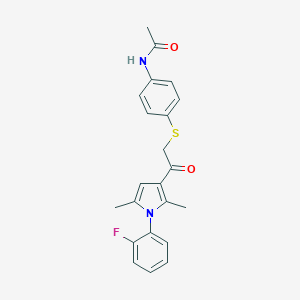 MLS000335684 MLS000335684 | C22H21FN2O2S | 396.48 | 4 / 1 | 4.3 | Yes |
| 206361 | 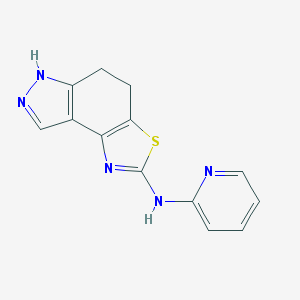 CHEMBL1830698 CHEMBL1830698 | C13H11N5S | 269.326 | 5 / 2 | 2.4 | Yes |
| 208547 | 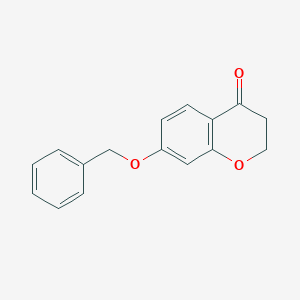 7-(benzyloxy)-2,3-dihydro-4H-chromen-4-one 7-(benzyloxy)-2,3-dihydro-4H-chromen-4-one | C16H14O3 | 254.285 | 3 / 0 | 2.8 | Yes |
| 489704 | 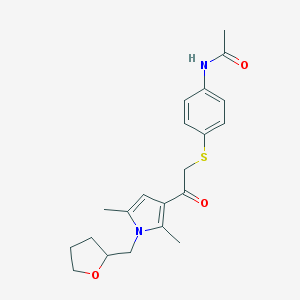 AC1N0VGZ AC1N0VGZ | C21H26N2O3S | 386.51 | 4 / 1 | 2.9 | Yes |
| 490016 | 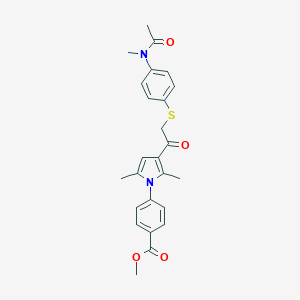 CHEMBL3561870 CHEMBL3561870 | C25H26N2O4S | 450.553 | 5 / 0 | 4.3 | Yes |
| 218676 | 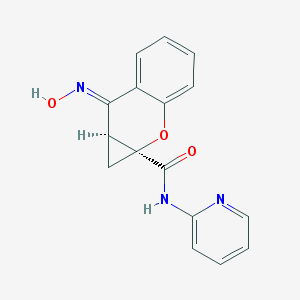 CID 70682300 CID 70682300 | C16H13N3O3 | 295.298 | 5 / 2 | 1.8 | Yes |
| 218947 | 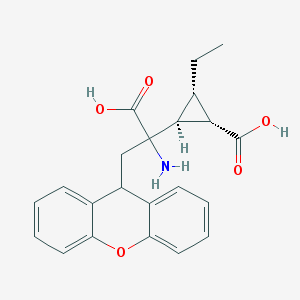 CHEMBL439775 CHEMBL439775 | C22H23NO5 | 381.428 | 6 / 3 | 0.7 | Yes |
| 221639 | 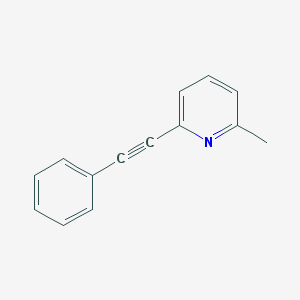 MPEP MPEP | C14H11N | 193.249 | 1 / 0 | 3.3 | Yes |
| 450594 | 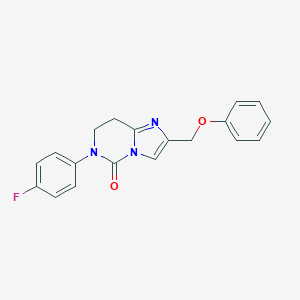 CHEMBL3401195 CHEMBL3401195 | C19H16FN3O2 | 337.354 | 4 / 0 | 3.1 | Yes |
| 235586 | 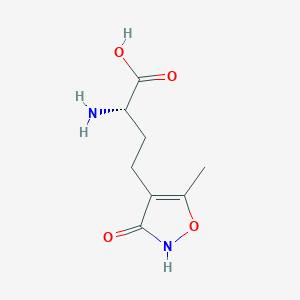 CHEMBL315268 CHEMBL315268 | C8H12N2O4 | 200.194 | 5 / 3 | -2.9 | Yes |
| 492626 | 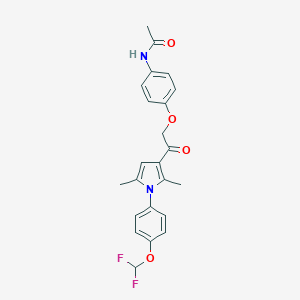 CHEMBL3560612 CHEMBL3560612 | C23H22F2N2O4 | 428.436 | 6 / 1 | 4.6 | Yes |
| 554537 | 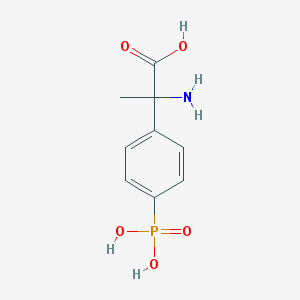 MPPG MPPG | C9H12NO5P | 245.171 | 6 / 4 | -3.5 | Yes |
| 255968 | 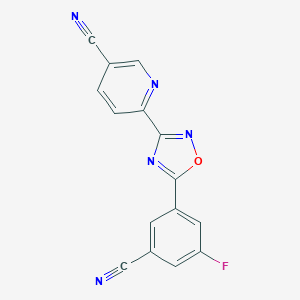 AZD-6538 AZD-6538 | C15H6FN5O | 291.245 | 7 / 0 | 2.0 | Yes |
| 529240 | 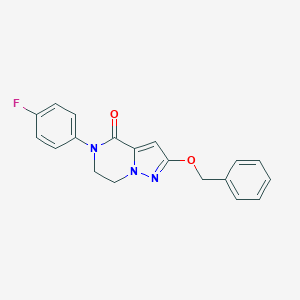 CHEMBL3747116 CHEMBL3747116 | C19H16FN3O2 | 337.354 | 4 / 0 | 3.2 | Yes |
| 273210 | 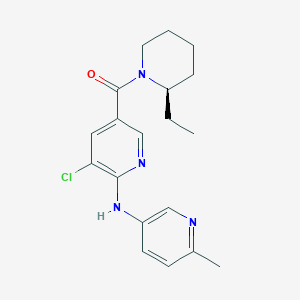 UNII-J5PMS063XX UNII-J5PMS063XX | C19H23ClN4O | 358.87 | 4 / 1 | 4.0 | Yes |
| 274093 | 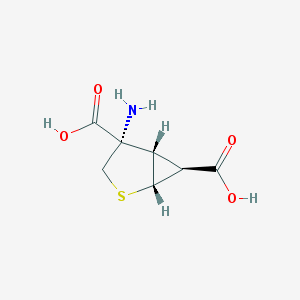 CHEMBL8839 CHEMBL8839 | C7H9NO4S | 203.212 | 6 / 3 | -3.5 | Yes |
| 274100 | 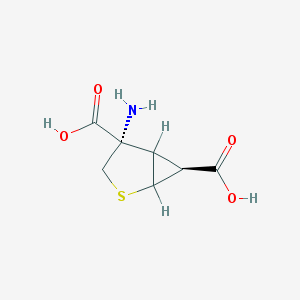 CHEMBL88999 CHEMBL88999 | C7H9NO4S | 203.212 | 6 / 3 | -3.5 | Yes |
| 497781 | 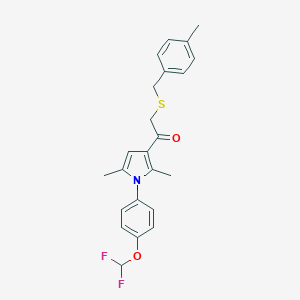 CHEMBL3560758 CHEMBL3560758 | C23H23F2NO2S | 415.499 | 5 / 0 | 6.3 | No |
| 529677 | 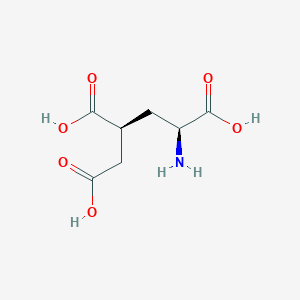 CHEMBL479797 CHEMBL479797 | C7H11NO6 | 205.166 | 7 / 4 | -4.0 | Yes |
| 290096 | 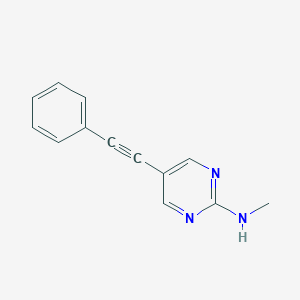 CHEMBL505105 CHEMBL505105 | C13H11N3 | 209.252 | 3 / 1 | 2.5 | Yes |
| 291056 | 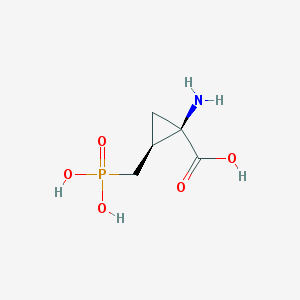 CHEMBL34880 CHEMBL34880 | C5H10NO5P | 195.111 | 6 / 4 | -5.0 | Yes |
| 291057 | 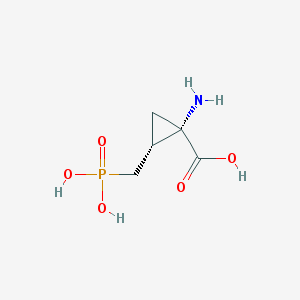 CHEMBL229697 CHEMBL229697 | C5H10NO5P | 195.111 | 6 / 4 | -5.0 | Yes |
| 291064 | 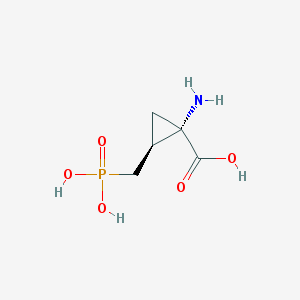 CHEMBL227288 CHEMBL227288 | C5H10NO5P | 195.111 | 6 / 4 | -5.0 | Yes |
| 291068 | 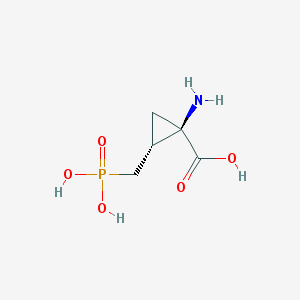 CHEMBL287703 CHEMBL287703 | C5H10NO5P | 195.111 | 6 / 4 | -5.0 | Yes |
| 292386 | 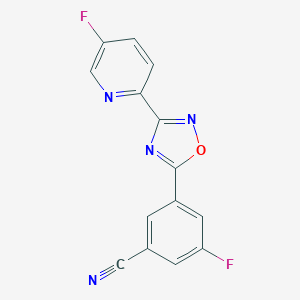 AZD 9272 AZD 9272 | C14H6F2N4O | 284.226 | 7 / 0 | 2.3 | Yes |
| 295457 | 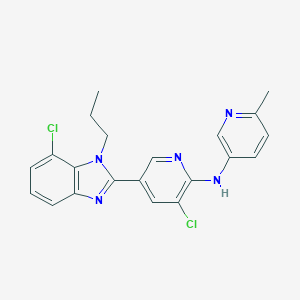 CHEMBL1672539 CHEMBL1672539 | C21H19Cl2N5 | 412.318 | 4 / 1 | 5.3 | No |
| 301609 | 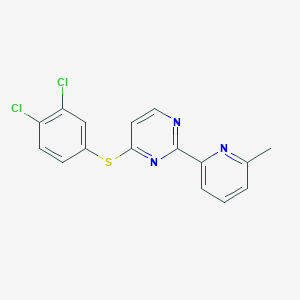 CHEMBL205571 CHEMBL205571 | C16H11Cl2N3S | 348.245 | 4 / 0 | 4.9 | Yes |
| 303022 | 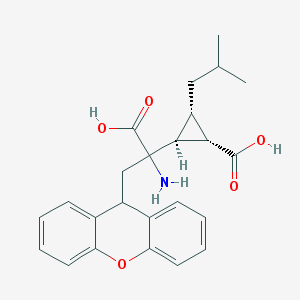 CHEMBL97200 CHEMBL97200 | C24H27NO5 | 409.482 | 6 / 3 | 1.5 | Yes |
| 305945 | 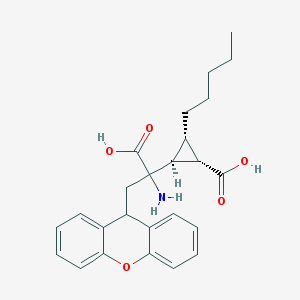 CHEMBL329920 CHEMBL329920 | C25H29NO5 | 423.509 | 6 / 3 | 2.4 | Yes |
| 500777 | 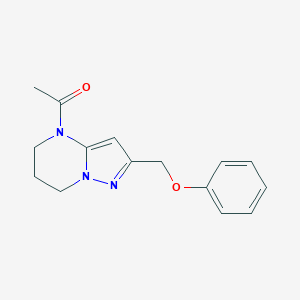 CHEMBL3633943 CHEMBL3633943 | C15H17N3O2 | 271.32 | 3 / 0 | 1.5 | Yes |
| 454047 | 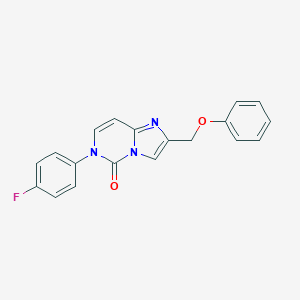 CHEMBL3401177 CHEMBL3401177 | C19H14FN3O2 | 335.338 | 4 / 0 | 3.2 | Yes |
| 315382 | 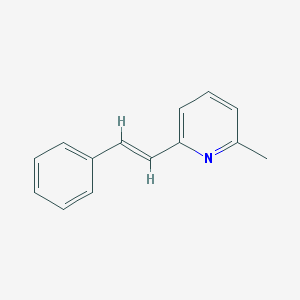 SIB 1893 SIB 1893 | C14H13N | 195.265 | 1 / 0 | 3.6 | Yes |
| 318414 | 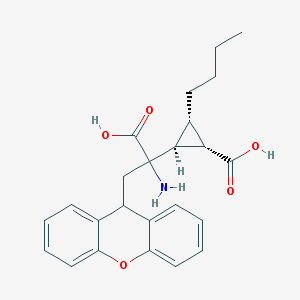 CHEMBL95868 CHEMBL95868 | C24H27NO5 | 409.482 | 6 / 3 | 1.8 | Yes |
| 324757 | 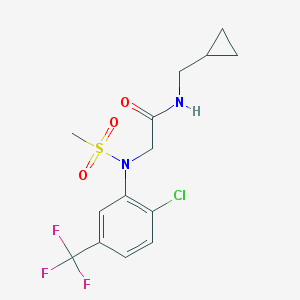 ML332 ML332 | C14H16ClF3N2O3S | 384.798 | 7 / 1 | 2.7 | Yes |
| 326892 | 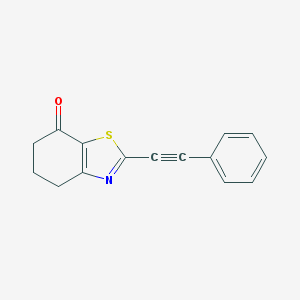 CHEMBL2426621 CHEMBL2426621 | C15H11NOS | 253.319 | 3 / 0 | 3.5 | Yes |
| 346322 | 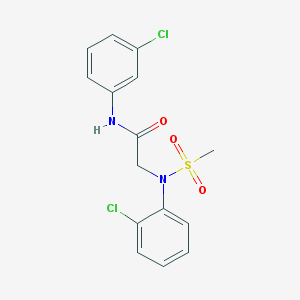 MLS000530551 MLS000530551 | C15H14Cl2N2O3S | 373.248 | 4 / 1 | 3.2 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218