

You can:
| Name | Glucagon receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Gcgr |
| Synonym | GGR GL-R glucagon receptor GR |
| Disease | N/A for non-human GPCRs |
| Length | 485 |
| Amino acid sequence | MLLTQLHCPYLLLLLVVLSCLPKAPSAQVMDFLFEKWKLYSDQCHHNLSLLPPPTELVCNRTFDKYSCWPDTPPNTTANISCPWYLPWYHKVQHRLVFKRCGPDGQWVRGPRGQSWRDASQCQMDDDEIEVQKGVAKMYSSYQVMYTVGYSLSLGALLLALVILLGLRKLHCTRNYIHGNLFASFVLKAGSVLVIDWLLKTRYSQKIGDDLSVSVWLSDGAVAGCRVATVIMQYGIIANYCWLLVEGVYLYSLLSITTFSEKSFFSLYLCIGWGSPLLFVIPWVVVKCLFENVQCWTSNDNMGFWWILRIPVLLAILINFFIFVRIIHLLVAKLRAHQMHYADYKFRLARSTLTLIPLLGVHEVVFAFVTDEHAQGTLRSTKLFFDLFFSSFQGLLVAVLYCFLNKEVQAELLRRWRRWQEGKALQEERMASSHGSHMAPAGTCHGDPCEKLQLMSAGSSSGTGCEPSAKTSLASSLPRLADSPT |
| UniProt | P30082 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4720 |
| IUPHAR | 251 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 1183 | 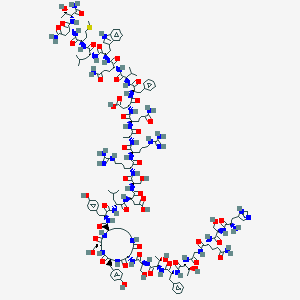 c[Asp9, Lys12,] glucagon-NH2 c[Asp9, Lys12,] glucagon-NH2 | C153H224N44O47S | 3463.8 | 52 / 54 | -13.7 | No |
| 5102 | 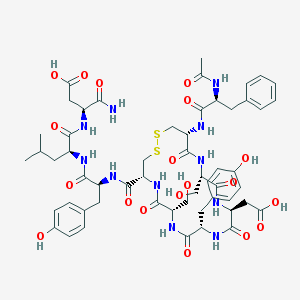 CHEMBL265503 CHEMBL265503 | C55H71N11O19S2 | 1254.35 | 21 / 17 | -1.0 | No |
| 6097 | 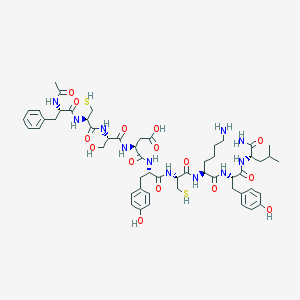 CHEMBL265501 CHEMBL265501 | C54H75N11O15S2 | 1182.38 | 18 / 17 | -2.4 | No |
| 9083 | 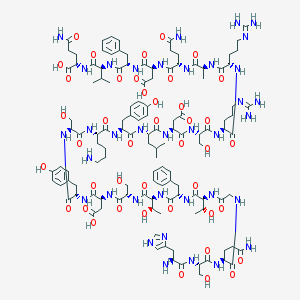 CHEMBL415177 CHEMBL415177 | C123H182N36O42 | 2837.02 | 47 / 45 | -18.0 | No |
| 11390 | 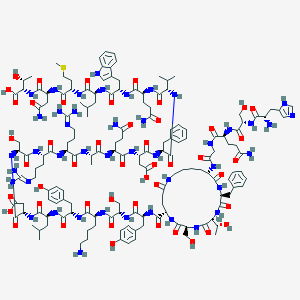 CID 77078079 CID 77078079 | C156H230N44O47S | 3505.88 | 53 / 52 | -15.4 | No |
| 442452 | 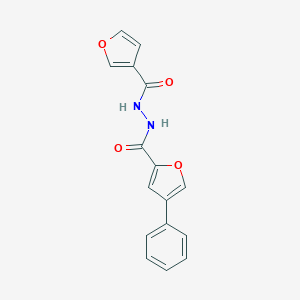 CHEMBL3359004 CHEMBL3359004 | C16H12N2O4 | 296.282 | 4 / 2 | 2.5 | Yes |
| 442942 | 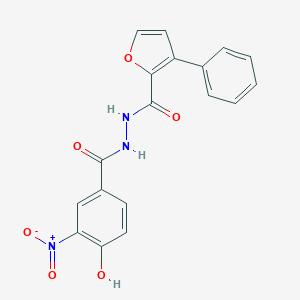 CHEMBL3326178 CHEMBL3326178 | C18H13N3O6 | 367.317 | 6 / 3 | 3.4 | Yes |
| 33360 | 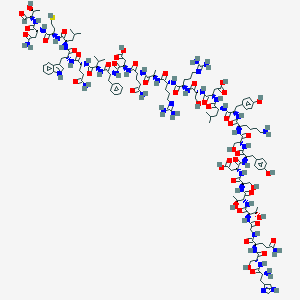 CID 44319559 CID 44319559 | C143H214N42O47S | 3305.59 | 53 / 52 | -19.5 | No |
| 36903 | 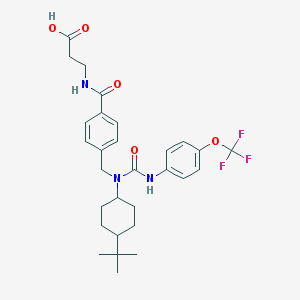 GRA EX-25 GRA EX-25 | C29H36F3N3O5 | 563.618 | 8 / 3 | 5.8 | No |
| 40485 | 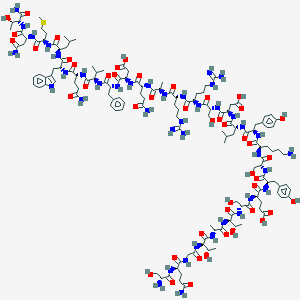 BDBM50059363 BDBM50059363 | C142H217N41O47S | 3282.6 | 52 / 53 | -18.3 | No |
| 443499 | 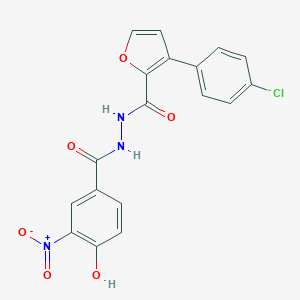 CHEMBL3326184 CHEMBL3326184 | C18H12ClN3O6 | 401.759 | 6 / 3 | 4.0 | Yes |
| 443558 | 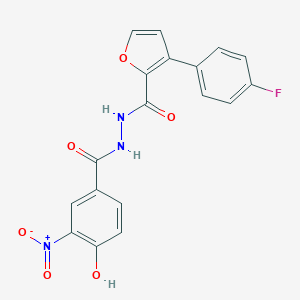 CHEMBL3326183 CHEMBL3326183 | C18H12FN3O6 | 385.307 | 7 / 3 | 3.5 | Yes |
| 49509 | 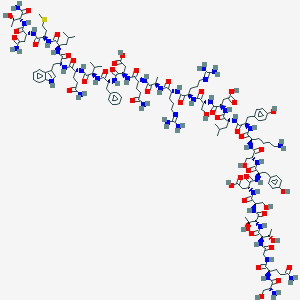 CID 44319561 CID 44319561 | C138H210N40O46S | 3197.49 | 51 / 50 | -19.2 | No |
| 50148 | 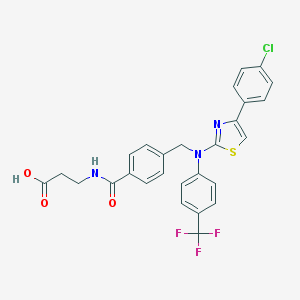 643011-22-7 643011-22-7 | C27H21ClF3N3O3S | 559.988 | 9 / 2 | 6.4 | No |
| 51813 | 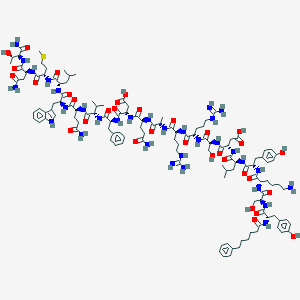 BDBM50098567 BDBM50098567 | C125H184N32O33S | 2695.1 | 37 / 39 | -5.0 | No |
| 56648 | 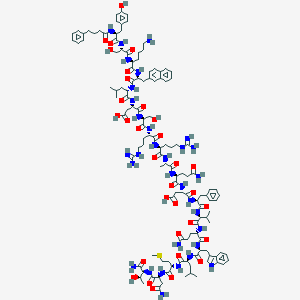 BDBM50098574 BDBM50098574 | C127H182N32O32S | 2701.1 | 36 / 38 | -4.5 | No |
| 59459 | 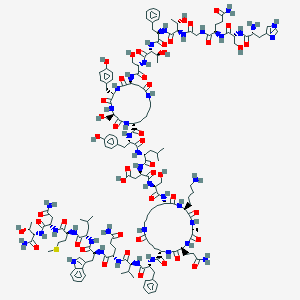 CID 44292844 CID 44292844 | C154H224N40O46S | 3403.78 | 50 / 49 | -11.2 | No |
| 60216 | 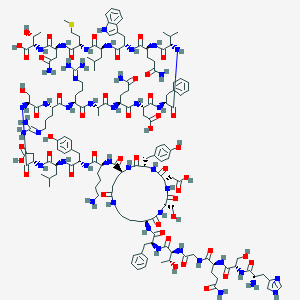 CID 44335736 CID 44335736 | C157H230N44O48S | 3533.89 | 54 / 52 | -15.8 | No |
| 64649 | 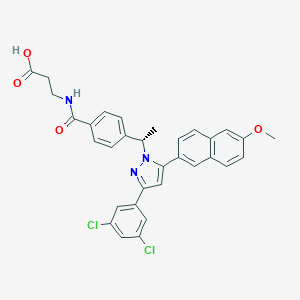 MK 0893 MK 0893 | C32H27Cl2N3O4 | 588.485 | 5 / 2 | 6.8 | No |
| 444300 | 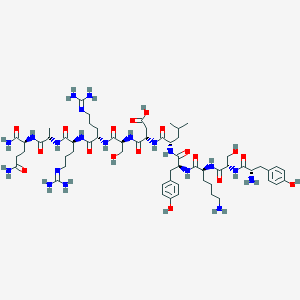 Glucagon(10-20)amide Glucagon(10-20)amide | C60H96N20O18 | 1385.55 | 22 / 23 | -9.2 | No |
| 444698 | 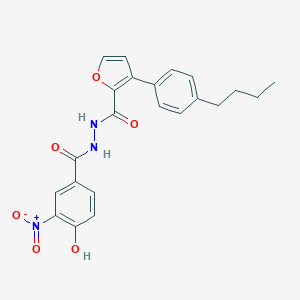 CHEMBL3326187 CHEMBL3326187 | C22H21N3O6 | 423.425 | 6 / 3 | 5.3 | No |
| 444804 | 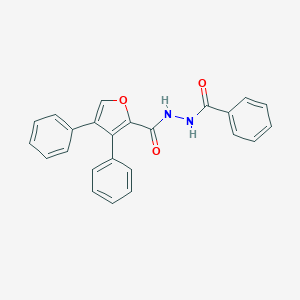 CHEMBL3326164 CHEMBL3326164 | C24H18N2O3 | 382.419 | 3 / 2 | 5.0 | Yes |
| 79817 | 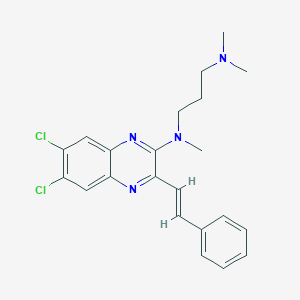 CHEMBL557974 CHEMBL557974 | C22H24Cl2N4 | 415.362 | 4 / 0 | 5.6 | No |
| 80050 | 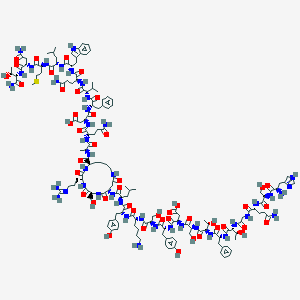 CHEMBL2372773 CHEMBL2372773 | C153H224N42O47S | 3435.78 | 52 / 51 | -16.4 | No |
| 80051 |  CHEMBL266412 CHEMBL266412 | C153H224N42O47S | 3435.78 | 52 / 51 | -16.4 | No |
| 83044 |  CID 44293064 CID 44293064 | C154H226N40O47S | 3421.79 | 52 / 50 | -15.6 | No |
| 560028 |  CHEMBL107070 CHEMBL107070 | C18H13ClN2O3 | 340.763 | 4 / 3 | 4.0 | Yes |
| 91067 |  CHEMBL442499 CHEMBL442499 | C150H222N42O47S | 3397.73 | 52 / 52 | -17.4 | No |
| 94570 | 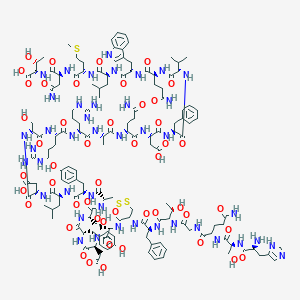 BDBM50104042 BDBM50104042 | C149H214N42O48S3 | 3457.78 | 55 / 53 | -13.0 | No |
| 445744 | 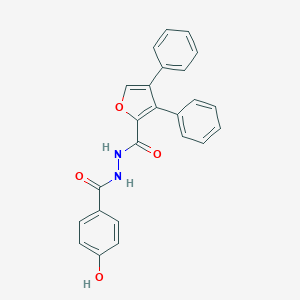 CHEMBL3326165 CHEMBL3326165 | C24H18N2O4 | 398.418 | 4 / 3 | 4.7 | Yes |
| 107111 | 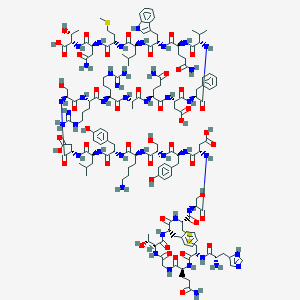 BDBM50104032 BDBM50104032 | C152H221N43O47S3 | 3498.87 | 55 / 53 | -15.0 | No |
| 108541 | 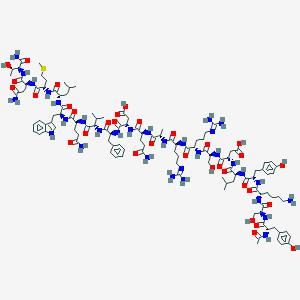 CID 44278164 CID 44278164 | C115H172N32O33S | 2562.89 | 37 / 37 | -9.1 | No |
| 109514 | 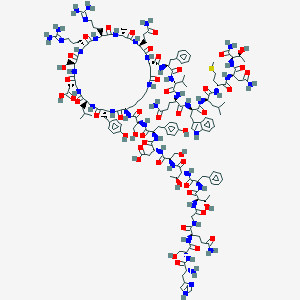 CID 44292665 CID 44292665 | C153H224N44O47S | 3463.8 | 52 / 52 | -14.5 | No |
| 109686 | 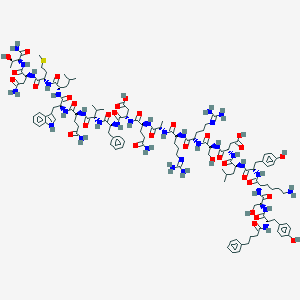 CID 44278221 CID 44278221 | C123H180N32O33S | 2667.04 | 37 / 37 | -6.9 | No |
| 115018 | 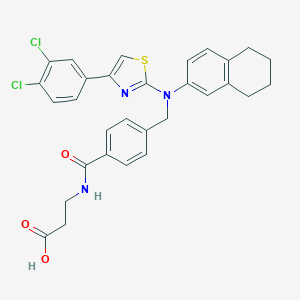 aminothiazole, 10 aminothiazole, 10 | C30H27Cl2N3O3S | 580.524 | 6 / 2 | 7.5 | No |
| 446236 | 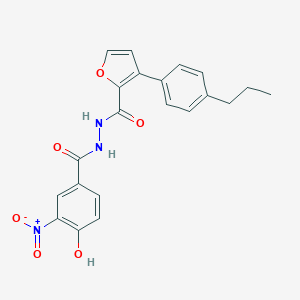 CHEMBL3325456 CHEMBL3325456 | C21H19N3O6 | 409.398 | 6 / 3 | 4.8 | Yes |
| 119646 | 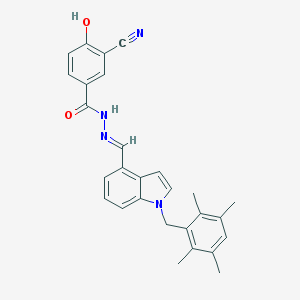 CHEMBL152640 CHEMBL152640 | C28H26N4O2 | 450.542 | 4 / 2 | 5.9 | No |
| 124826 | 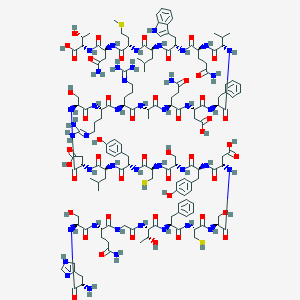 CID 44335779 CID 44335779 | C149H216N42O48S3 | 3459.79 | 55 / 53 | -14.7 | No |
| 446620 | 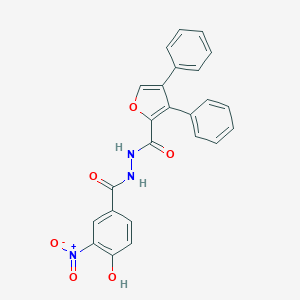 CHEMBL3326175 CHEMBL3326175 | C24H17N3O6 | 443.415 | 6 / 3 | 5.0 | Yes |
| 127975 | 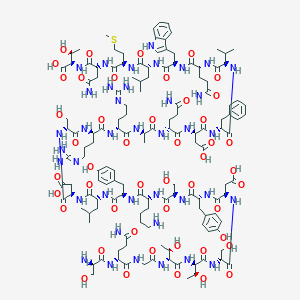 CID 44319558 CID 44319558 | C138H209N39O47S | 3198.48 | 52 / 50 | -18.5 | No |
| 128718 | 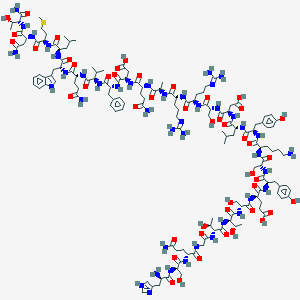 CHEMBL412352 CHEMBL412352 | C145H219N43O47S | 3348.66 | 53 / 52 | -19.3 | No |
| 129011 | 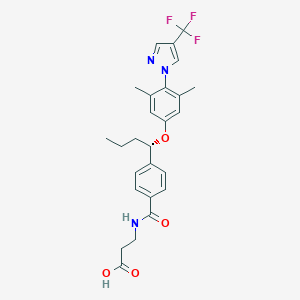 glucagon receptor antagonists-4 glucagon receptor antagonists-4 | C26H28F3N3O4 | 503.522 | 8 / 2 | 5.0 | No |
| 132421 | 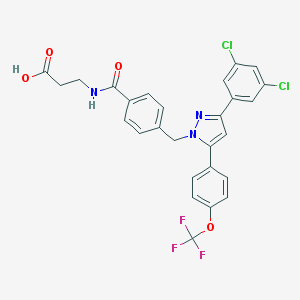 CHEMBL1644183 CHEMBL1644183 | C27H20Cl2F3N3O4 | 578.369 | 8 / 2 | 6.4 | No |
| 139156 | 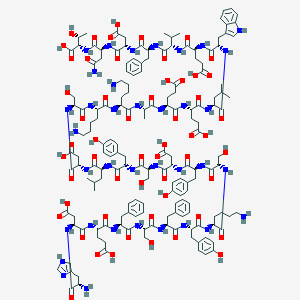 CHEMBL268583 CHEMBL268583 | C165H226N36O55 | 3593.82 | 60 / 52 | -15.9 | No |
| 447226 | 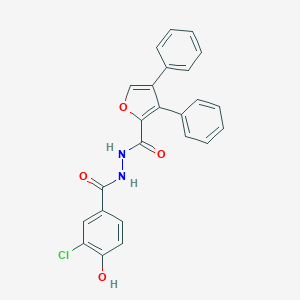 CHEMBL3326172 CHEMBL3326172 | C24H17ClN2O4 | 432.86 | 4 / 3 | 5.3 | No |
| 447813 | 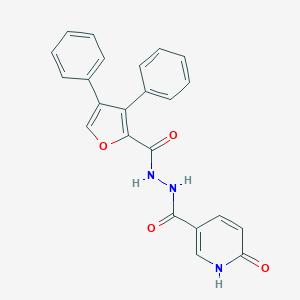 CHEMBL3326170 CHEMBL3326170 | C23H17N3O4 | 399.406 | 4 / 3 | 3.0 | Yes |
| 448322 | 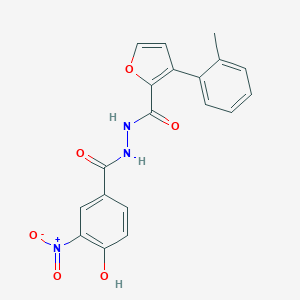 CHEMBL3326179 CHEMBL3326179 | C19H15N3O6 | 381.344 | 6 / 3 | 3.8 | Yes |
| 174426 | 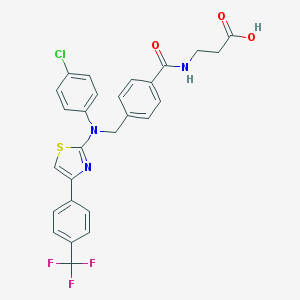 aminothiazole, 16 aminothiazole, 16 | C27H21ClF3N3O3S | 559.988 | 9 / 2 | 6.4 | No |
| 175218 | 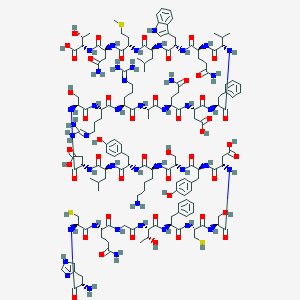 CID 44335967 CID 44335967 | C152H223N43O47S3 | 3500.89 | 55 / 53 | -16.2 | No |
| 448706 | 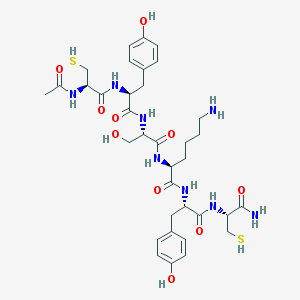 CHEMBL28541 CHEMBL28541 | C35H50N8O10S2 | 806.951 | 13 / 13 | -1.4 | No |
| 179765 | 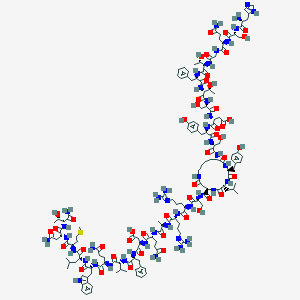 CID 77077841 CID 77077841 | C153H224N44O47S | 3463.8 | 52 / 52 | -15.1 | No |
| 448940 | 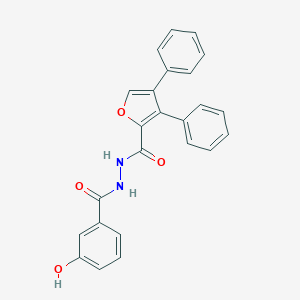 CHEMBL3326166 CHEMBL3326166 | C24H18N2O4 | 398.418 | 4 / 3 | 4.7 | Yes |
| 449233 | 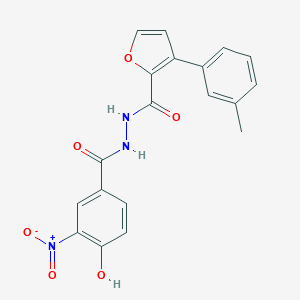 CHEMBL3326180 CHEMBL3326180 | C19H15N3O6 | 381.344 | 6 / 3 | 3.8 | Yes |
| 449540 | 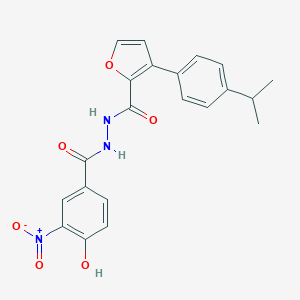 CHEMBL3326186 CHEMBL3326186 | C21H19N3O6 | 409.398 | 6 / 3 | 4.5 | Yes |
| 200982 | 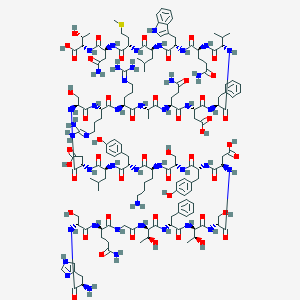 CHEMBL429362 CHEMBL429362 | C153H225N43O49S | 3482.79 | 55 / 53 | -17.7 | No |
| 200983 | 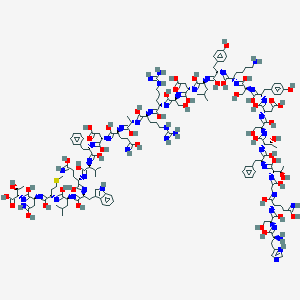 CID 102331734 CID 102331734 | C153H225N43O49S | 3482.79 | 87 / 59 | 0.8 | No |
| 201324 | 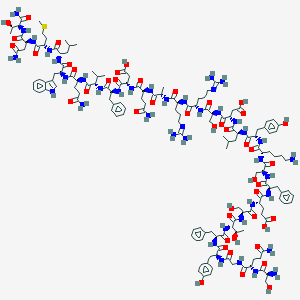 CID 71456274 CID 71456274 | C153H223N41O46S | 3404.77 | 51 / 50 | -15.3 | No |
| 201325 | 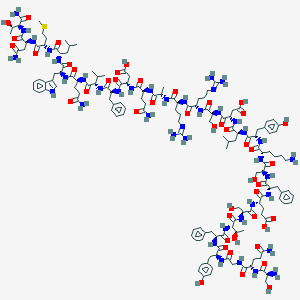 CID 71450989 CID 71450989 | C153H223N41O46S | 3404.77 | 51 / 50 | -15.3 | No |
| 449738 | 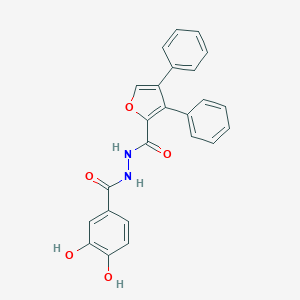 SCHEMBL3982429 SCHEMBL3982429 | C24H18N2O5 | 414.417 | 5 / 4 | 4.3 | Yes |
| 450585 | 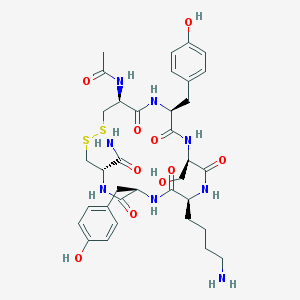 CHEMBL282363 CHEMBL282363 | C35H48N8O10S2 | 804.935 | 13 / 11 | -0.9 | No |
| 227076 |  CID 44278463 CID 44278463 | C121H176N32O33S | 2638.99 | 37 / 37 | -7.5 | No |
| 230325 |  CHEMBL411904 CHEMBL411904 | C86H127N25O30 | 1991.11 | 34 / 33 | -15.0 | No |
| 239428 |  CHEMBL442492 CHEMBL442492 | C165H224N36O49 | 3495.81 | 55 / 50 | -11.0 | No |
| 257501 |  CHEMBL216417 CHEMBL216417 | C51H68N10O16S2 | 1141.28 | 18 / 17 | -0.9 | No |
| 259058 |  CID 77078086 CID 77078086 | C150H219N43O49S3 | 3504.83 | 57 / 55 | -19.2 | No |
| 452110 |  CHEMBL3326171 CHEMBL3326171 | C24H17FN2O4 | 416.408 | 5 / 3 | 4.8 | Yes |
| 265461 |  CID 44336110 CID 44336110 | C154H225N41O48S | 3450.79 | 53 / 51 | -15.4 | No |
| 267610 |  CID 71458083 CID 71458083 | C147H219N41O46S | 3328.67 | 51 / 50 | -16.9 | No |
| 267611 | 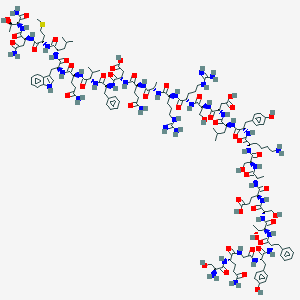 BDBM50407841 BDBM50407841 | C147H219N41O46S | 3328.67 | 51 / 52 | -16.1 | No |
| 452516 | 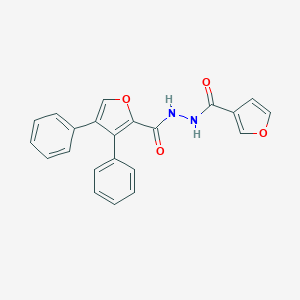 SCHEMBL3976454 SCHEMBL3976454 | C22H16N2O4 | 372.38 | 4 / 2 | 4.1 | Yes |
| 452541 | 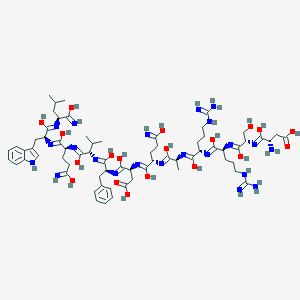 Glucagon(15-26)amide Glucagon(15-26)amide | C67H102N22O19 | 1519.69 | 36 / 28 | 0.4 | No |
| 452880 | 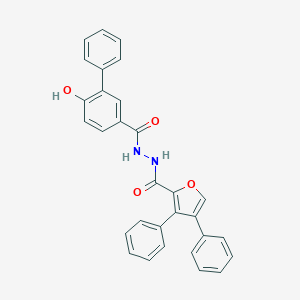 CHEMBL3326177 CHEMBL3326177 | C30H22N2O4 | 474.516 | 4 / 3 | 6.3 | No |
| 292656 | 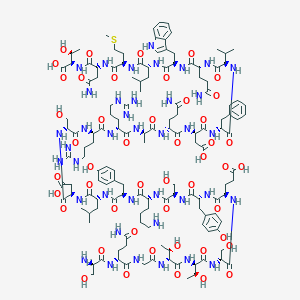 [des-Phe6, Glu9] glucagon [des-Phe6, Glu9] glucagon | C139H211N39O47S | 3212.5 | 52 / 52 | -17.4 | No |
| 306505 | 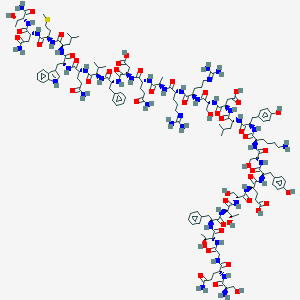 CHEMBL2369990 CHEMBL2369990 | C148H221N41O47S | 3358.7 | 52 / 51 | -17.5 | No |
| 306506 | 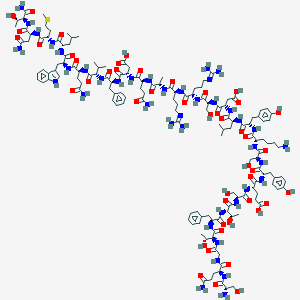 CID 44278168 CID 44278168 | C148H221N41O47S | 3358.7 | 52 / 51 | -17.5 | No |
| 307246 | 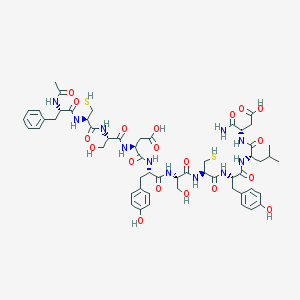 CHEMBL267646 CHEMBL267646 | C55H73N11O19S2 | 1256.37 | 21 / 19 | -2.0 | No |
| 454184 | 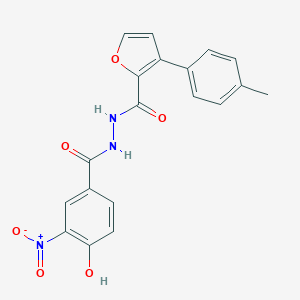 CHEMBL3326181 CHEMBL3326181 | C19H15N3O6 | 381.344 | 6 / 3 | 3.8 | Yes |
| 314519 | 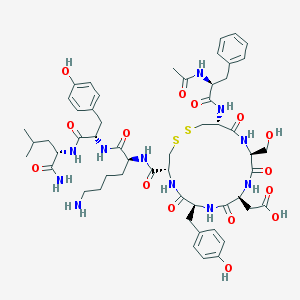 CHEMBL408316 CHEMBL408316 | C54H73N11O15S2 | 1180.36 | 18 / 15 | -1.9 | No |
| 454194 | 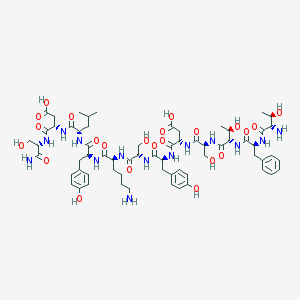 Glucagon(5-16)amide Glucagon(5-16)amide | C64H92N14O23 | 1425.52 | 25 / 23 | -9.7 | No |
| 318386 | 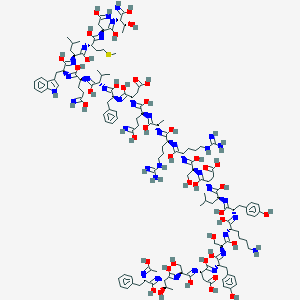 Ac-glucagon(6-29)amide Ac-glucagon(6-29)amide | C135H198N36O41S | 3013.34 | 73 / 50 | 5.3 | No |
| 318394 |  CHEMBL413666 CHEMBL413666 | C153H223N41O47S | 3420.77 | 52 / 51 | -15.7 | No |
| 318395 |  CHEMBL2369127 CHEMBL2369127 | C153H223N41O47S | 3420.77 | 52 / 51 | -15.7 | No |
| 454515 |  CHEMBL3326185 CHEMBL3326185 | C20H17N3O6 | 395.371 | 6 / 3 | 4.2 | Yes |
| 323891 |  CID 44278464 CID 44278464 | C123H176N32O31S | 2631.01 | 35 / 35 | -5.9 | No |
| 324259 | 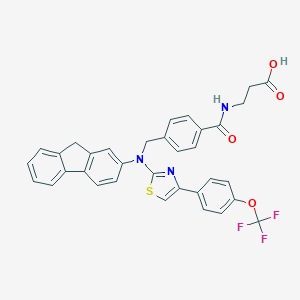 aminothiazole, 6 aminothiazole, 6 | C34H26F3N3O4S | 629.654 | 10 / 2 | 7.7 | No |
| 331386 | 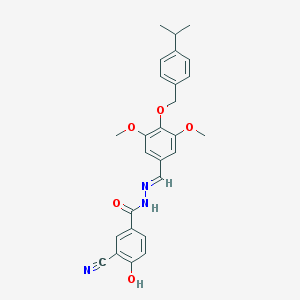 CHEMBL152765 CHEMBL152765 | C27H27N3O5 | 473.529 | 7 / 2 | 5.3 | No |
| 337006 | 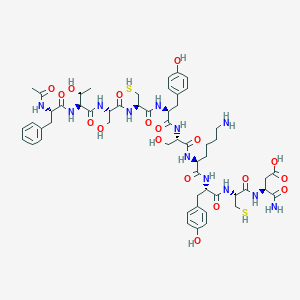 CHEMBL266166 CHEMBL266166 | C55H76N12O18S2 | 1257.4 | 21 / 20 | -5.6 | No |
| 343081 | 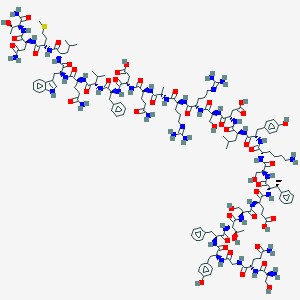 CHEMBL3037804 CHEMBL3037804 | C154H225N41O46S | 3418.79 | 51 / 50 | -15.0 | No |
| 343383 | 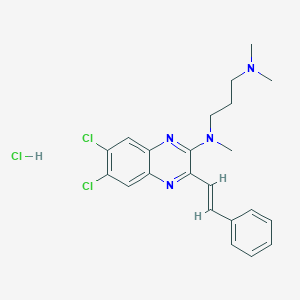 CHEMBL557974 CHEMBL557974 | C22H25Cl3N4 | 451.82 | 4 / 1 | N/A | N/A |
| 455387 | 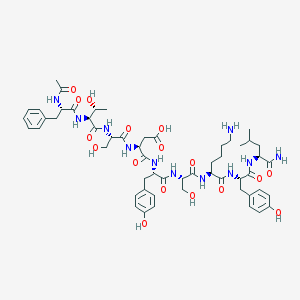 Ac-glucagon(6-14)amide Ac-glucagon(6-14)amide | C55H77N11O17 | 1164.28 | 18 / 17 | -3.9 | No |
| 344742 | 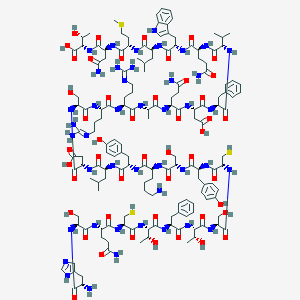 CID 77078081 CID 77078081 | C153H227N43O47S3 | 3516.93 | 55 / 54 | -16.6 | No |
| 347548 | 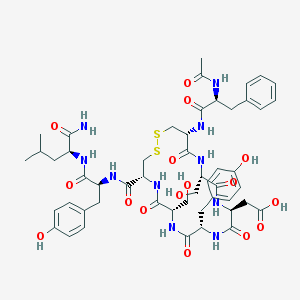 CHEMBL412578 CHEMBL412578 | C51H66N10O16S2 | 1139.26 | 18 / 15 | 0.1 | No |
| 347712 | 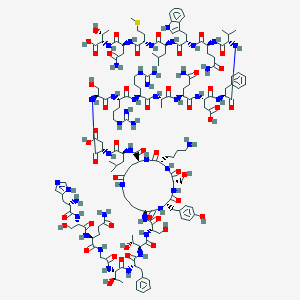 BDBM50104035 BDBM50104035 | C151H228N44O47S | 3443.81 | 53 / 54 | -16.5 | No |
| 350839 | 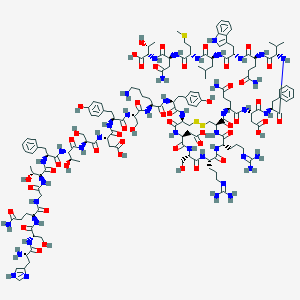 CID 44335807 CID 44335807 | C150H217N43O49S3 | 3502.82 | 57 / 53 | -18.7 | No |
| 352497 | 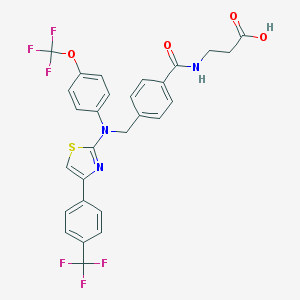 aminothiazole, 19 aminothiazole, 19 | C28H21F6N3O4S | 609.543 | 13 / 2 | 7.0 | No |
| 352717 | 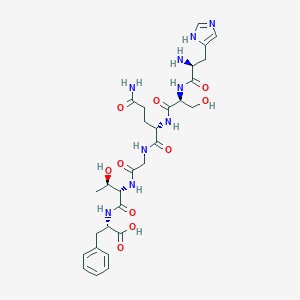 glucagon 1-6 glucagon 1-6 | C29H41N9O10 | 675.7 | 12 / 11 | -6.1 | No |
| 455800 | 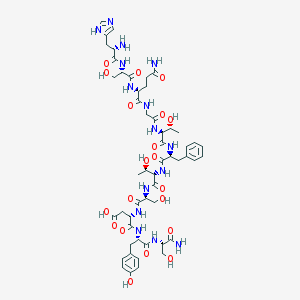 Glucagon(1-11)amide Glucagon(1-11)amide | C52H73N15O20 | 1228.24 | 22 / 21 | -10.4 | No |
| 355580 | 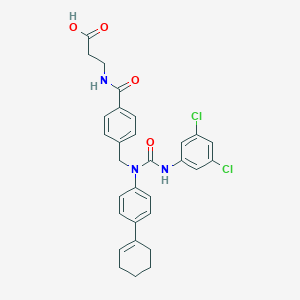 CHEMBL386446 CHEMBL386446 | C30H29Cl2N3O4 | 566.479 | 4 / 3 | 5.9 | No |
| 361984 | 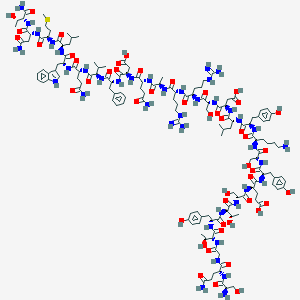 BDBM50059358 BDBM50059358 | C148H221N41O48S | 3374.7 | 53 / 54 | -17.1 | No |
| 366162 | 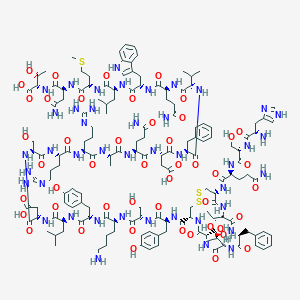 CID 44336094 CID 44336094 | C153H225N43O47S3 | 3514.92 | 55 / 52 | -15.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218