

You can:
| Name | Neuropeptides B/W receptor type 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | NPBWR1 |
| Synonym | G protein-coupled receptor 7 G-protein coupled receptor 7 GPR7 neuropeptides B/W receptor type 1 NPBW1 receptor |
| Disease | N/A |
| Length | 328 |
| Amino acid sequence | MDNASFSEPWPANASGPDPALSCSNASTLAPLPAPLAVAVPVVYAVICAVGLAGNSAVLYVLLRAPRMKTVTNLFILNLAIADELFTLVLPINIADFLLRQWPFGELMCKLIVAIDQYNTFSSLYFLTVMSADRYLVVLATAESRRVAGRTYSAARAVSLAVWGIVTLVVLPFAVFARLDDEQGRRQCVLVFPQPEAFWWRASRLYTLVLGFAIPVSTICVLYTTLLCRLHAMRLDSHAKALERAKKRVTFLVVAILAVCLLCWTPYHLSTVVALTTDLPQTPLVIAISYFITSLSYANSCLNPFLYAFLDASFRRNLRQLITCRAAA |
| UniProt | P48145 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P48145 |
| 3D structure model | This predicted structure model is from GPCR-EXP P48145. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1293293 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 2154 | 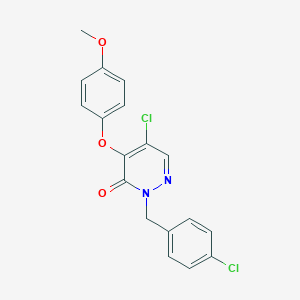 CHEMBL2314316 CHEMBL2314316 | C18H14Cl2N2O3 | 377.221 | 4 / 0 | 4.1 | Yes |
| 3891 | 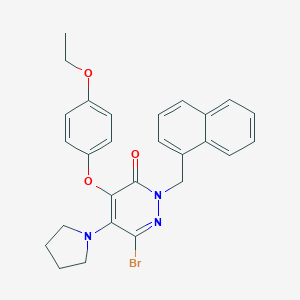 CHEMBL1995837 CHEMBL1995837 | C27H26BrN3O3 | 520.427 | 5 / 0 | 6.2 | No |
| 4163 | 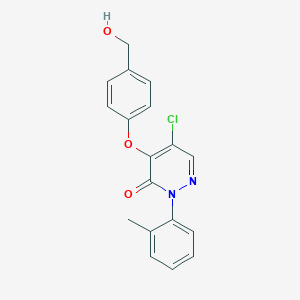 CHEMBL2314289 CHEMBL2314289 | C18H15ClN2O3 | 342.779 | 4 / 1 | 3.1 | Yes |
| 4559 | 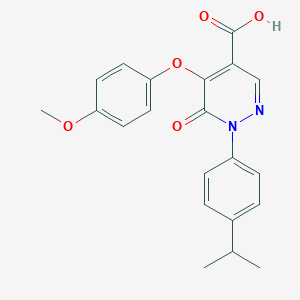 CHEMBL2207521 CHEMBL2207521 | C21H20N2O5 | 380.4 | 6 / 1 | 3.6 | Yes |
| 4815 | 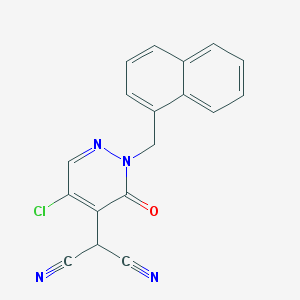 CHEMBL2001833 CHEMBL2001833 | C18H11ClN4O | 334.763 | 4 / 0 | 2.7 | Yes |
| 5358 | 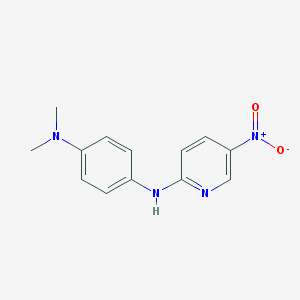 BAS 03293908 BAS 03293908 | C13H14N4O2 | 258.281 | 5 / 1 | 2.7 | Yes |
| 6060 | 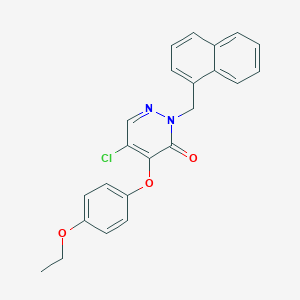 CHEMBL2314291 CHEMBL2314291 | C23H19ClN2O3 | 406.866 | 4 / 0 | 5.1 | No |
| 6353 | 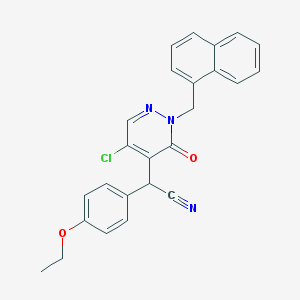 CHEMBL1973528 CHEMBL1973528 | C25H20ClN3O2 | 429.904 | 4 / 0 | 4.7 | Yes |
| 6704 | 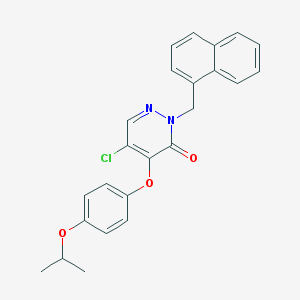 CHEMBL2314295 CHEMBL2314295 | C24H21ClN2O3 | 420.893 | 4 / 0 | 5.6 | No |
| 6794 | 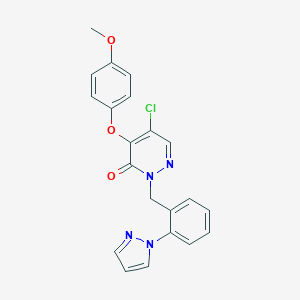 CHEMBL1981304 CHEMBL1981304 | C21H17ClN4O3 | 408.842 | 5 / 0 | 3.5 | Yes |
| 6957 | 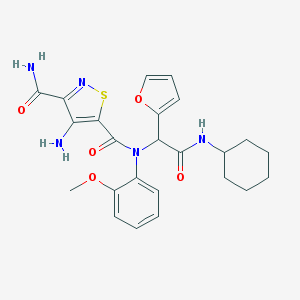 MLS001225295 MLS001225295 | C24H27N5O5S | 497.57 | 8 / 3 | 3.3 | Yes |
| 7714 | 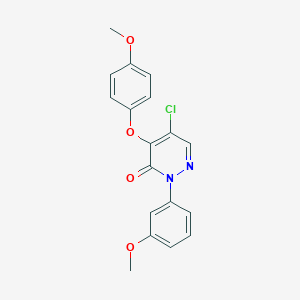 SR-02000000483 SR-02000000483 | C18H15ClN2O4 | 358.778 | 5 / 0 | 3.5 | Yes |
| 7888 | 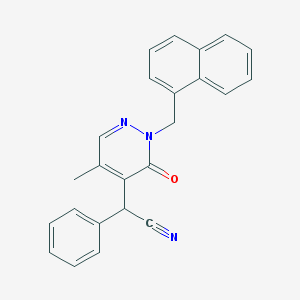 CHEMBL2003269 CHEMBL2003269 | C24H19N3O | 365.436 | 3 / 0 | 3.9 | Yes |
| 9076 | 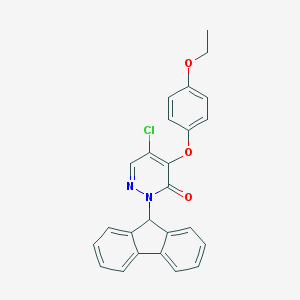 CHEMBL2314300 CHEMBL2314300 | C25H19ClN2O3 | 430.888 | 4 / 0 | 5.3 | No |
| 9305 | 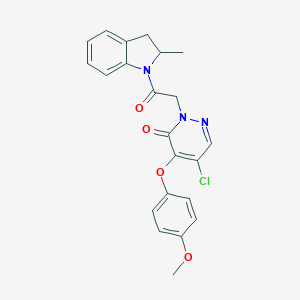 CHEMBL1968640 CHEMBL1968640 | C22H20ClN3O4 | 425.869 | 5 / 0 | 3.6 | Yes |
| 10421 | 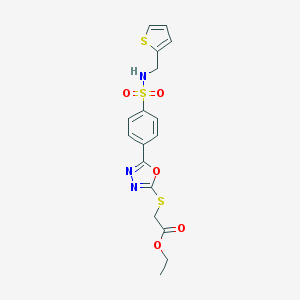 MLS000123364 MLS000123364 | C17H17N3O5S3 | 439.519 | 10 / 1 | 2.7 | Yes |
| 11965 | 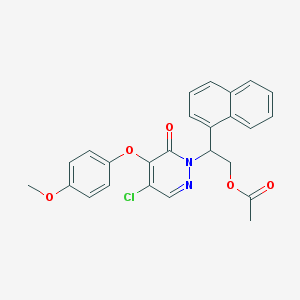 CHEMBL2314330 CHEMBL2314330 | C25H21ClN2O5 | 464.902 | 6 / 0 | 4.7 | Yes |
| 12603 | 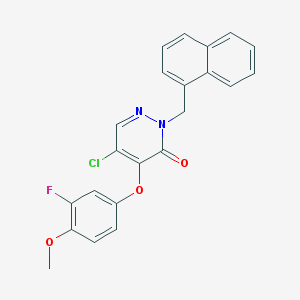 CHEMBL2005648 CHEMBL2005648 | C22H16ClFN2O3 | 410.829 | 5 / 0 | 4.9 | Yes |
| 14046 | 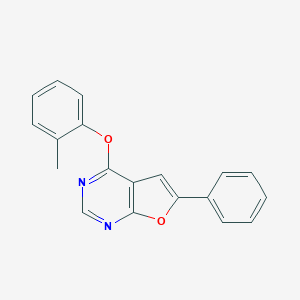 4-(2-methylphenoxy)-6-phenylfuro[2,3-d]pyrimidine 4-(2-methylphenoxy)-6-phenylfuro[2,3-d]pyrimidine | C19H14N2O2 | 302.333 | 4 / 0 | 4.8 | Yes |
| 14789 | 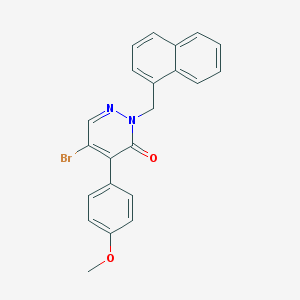 CHEMBL1991452 CHEMBL1991452 | C22H17BrN2O2 | 421.294 | 3 / 0 | 4.7 | Yes |
| 15404 | 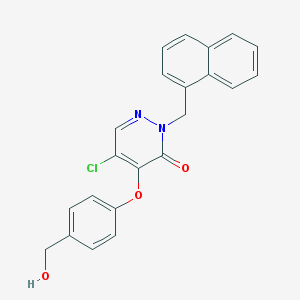 CHEMBL2314290 CHEMBL2314290 | C22H17ClN2O3 | 392.839 | 4 / 1 | 3.9 | Yes |
| 15654 | 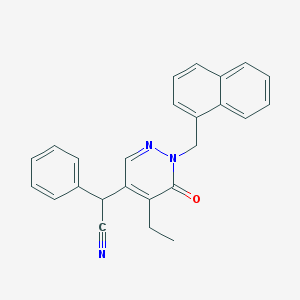 CHEMBL1985996 CHEMBL1985996 | C25H21N3O | 379.463 | 3 / 0 | 4.4 | Yes |
| 15915 | 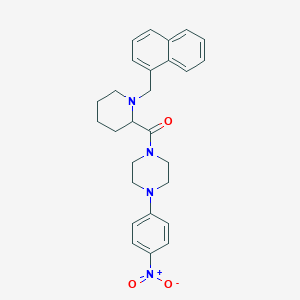 CHEMBL2068743 CHEMBL2068743 | C27H30N4O3 | 458.562 | 5 / 0 | 4.8 | Yes |
| 16993 | 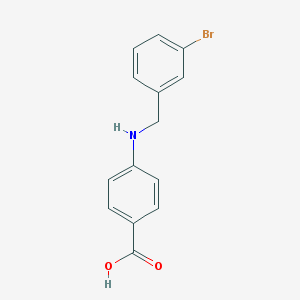 4-[(3-bromobenzyl)amino]benzoic acid 4-[(3-bromobenzyl)amino]benzoic acid | C14H12BrNO2 | 306.159 | 3 / 2 | 3.6 | Yes |
| 18178 | 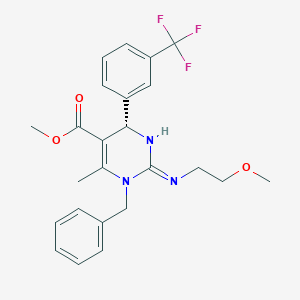 MLS000830374 MLS000830374 | C24H26F3N3O3 | 461.485 | 7 / 1 | 3.5 | Yes |
| 18725 | 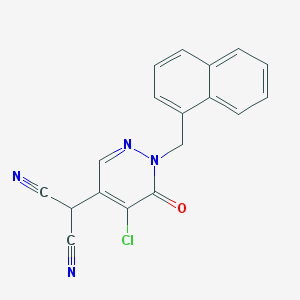 CHEMBL1972164 CHEMBL1972164 | C18H11ClN4O | 334.763 | 4 / 0 | 2.7 | Yes |
| 18914 | 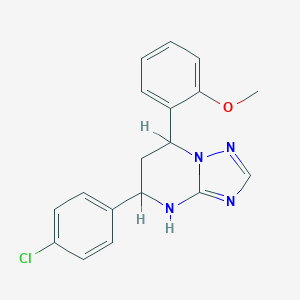 SMR000074933 SMR000074933 | C18H17ClN4O | 340.811 | 4 / 1 | 4.2 | Yes |
| 19743 | 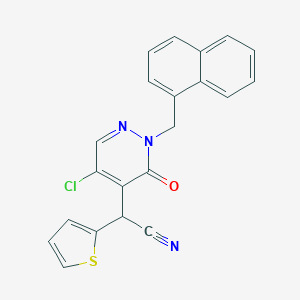 CHEMBL1977778 CHEMBL1977778 | C21H14ClN3OS | 391.873 | 4 / 0 | 4.1 | Yes |
| 21588 | 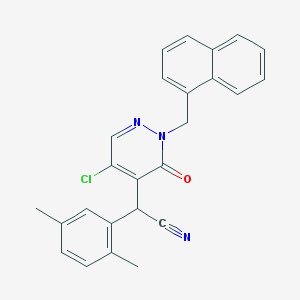 CHEMBL1974066 CHEMBL1974066 | C25H20ClN3O | 413.905 | 3 / 0 | 5.1 | No |
| 22768 | 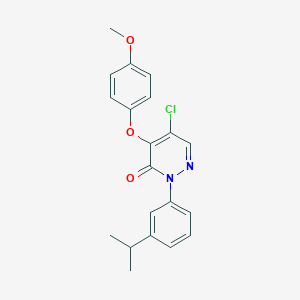 SR-02000000371 SR-02000000371 | C20H19ClN2O3 | 370.833 | 4 / 0 | 4.7 | Yes |
| 23392 | 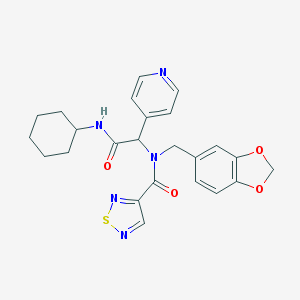 SMR000117962 SMR000117962 | C24H25N5O4S | 479.555 | 8 / 1 | 3.2 | Yes |
| 23490 | 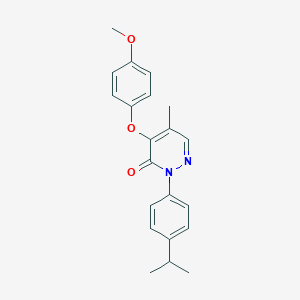 SR-02000000428 SR-02000000428 | C21H22N2O3 | 350.418 | 4 / 0 | 4.2 | Yes |
| 23663 | 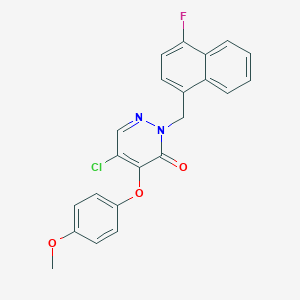 CHEMBL2314322 CHEMBL2314322 | C22H16ClFN2O3 | 410.829 | 5 / 0 | 4.9 | Yes |
| 24447 | 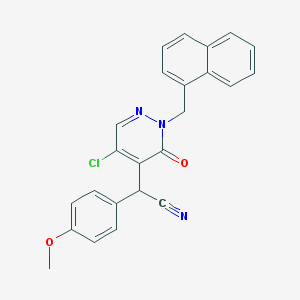 CHEMBL2002041 CHEMBL2002041 | C24H18ClN3O2 | 415.877 | 4 / 0 | 4.4 | Yes |
| 25897 | 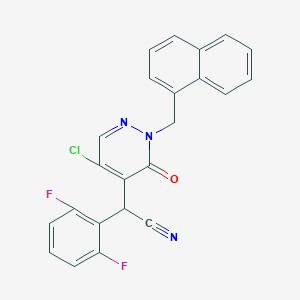 CHEMBL1990398 CHEMBL1990398 | C23H14ClF2N3O | 421.832 | 5 / 0 | 4.6 | Yes |
| 26104 | 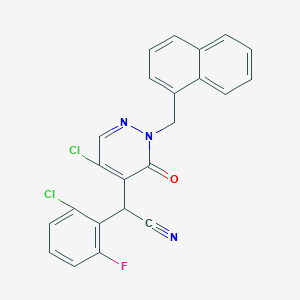 CHEMBL1979108 CHEMBL1979108 | C23H14Cl2FN3O | 438.283 | 4 / 0 | 5.1 | No |
| 26188 | 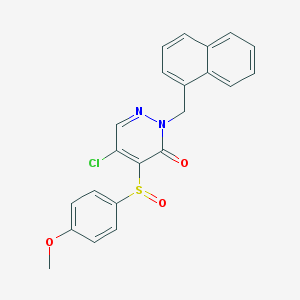 CHEMBL1987120 CHEMBL1987120 | C22H17ClN2O3S | 424.899 | 5 / 0 | 4.0 | Yes |
| 26336 | 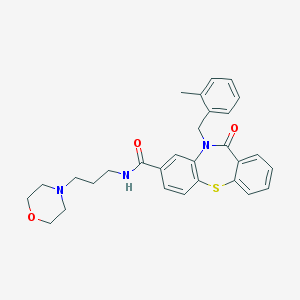 MLS001030841 MLS001030841 | C29H31N3O3S | 501.645 | 5 / 1 | 4.2 | No |
| 27022 | 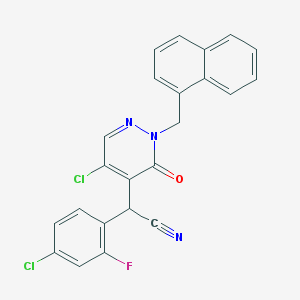 CHEMBL2001516 CHEMBL2001516 | C23H14Cl2FN3O | 438.283 | 4 / 0 | 5.1 | No |
| 28055 | 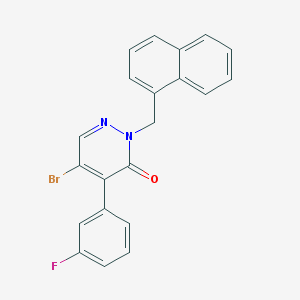 CHEMBL1998041 CHEMBL1998041 | C21H14BrFN2O | 409.258 | 3 / 0 | 4.9 | Yes |
| 28583 | 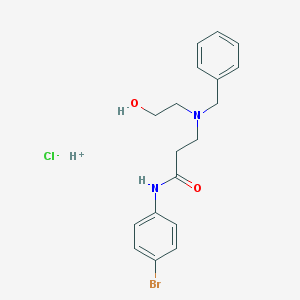 AC1LD3DJ AC1LD3DJ | C18H22BrClN2O2 | 413.74 | 4 / 3 | N/A | N/A |
| 29103 | 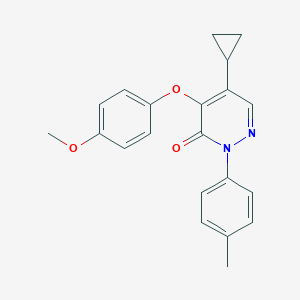 SR-02000000358 SR-02000000358 | C21H20N2O3 | 348.402 | 4 / 0 | 4.0 | Yes |
| 29770 | 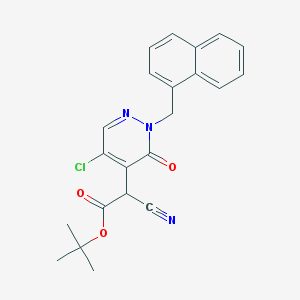 CHEMBL1999105 CHEMBL1999105 | C22H20ClN3O3 | 409.87 | 5 / 0 | 3.8 | Yes |
| 31010 | 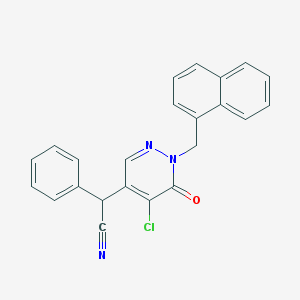 CHEMBL1966941 CHEMBL1966941 | C23H16ClN3O | 385.851 | 3 / 0 | 4.4 | Yes |
| 31309 | 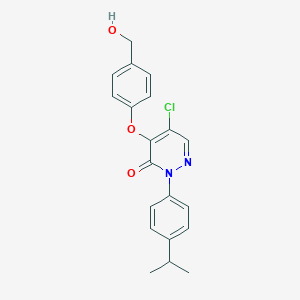 SR-02000000355 SR-02000000355 | C20H19ClN2O3 | 370.833 | 4 / 1 | 3.8 | Yes |
| 31394 | 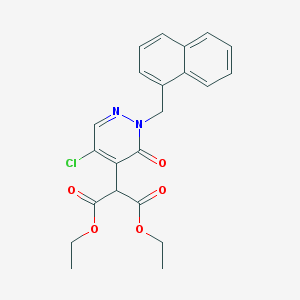 CHEMBL1989216 CHEMBL1989216 | C22H21ClN2O5 | 428.869 | 6 / 0 | 3.7 | Yes |
| 32692 | 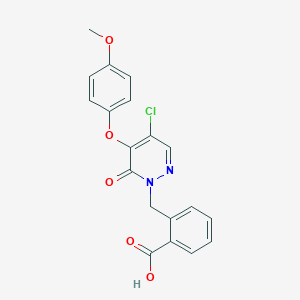 CHEMBL1965555 CHEMBL1965555 | C19H15ClN2O5 | 386.788 | 6 / 1 | 3.0 | Yes |
| 32881 | 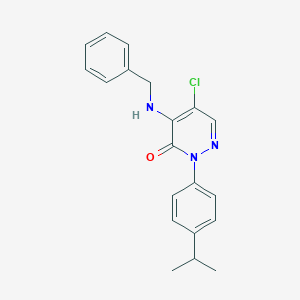 SR-02000000443 SR-02000000443 | C20H20ClN3O | 353.85 | 3 / 1 | 4.7 | Yes |
| 33426 | 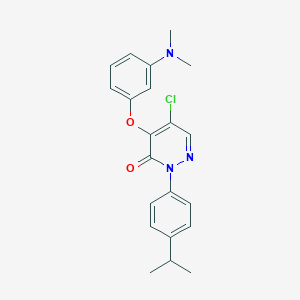 SR-02000000459 SR-02000000459 | C21H22ClN3O2 | 383.876 | 4 / 0 | 4.9 | Yes |
| 34389 | 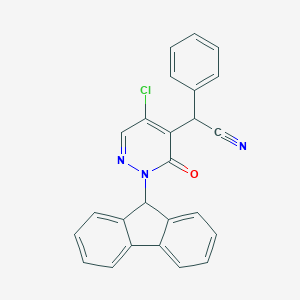 CHEMBL1990764 CHEMBL1990764 | C25H16ClN3O | 409.873 | 3 / 0 | 4.6 | Yes |
| 34691 | 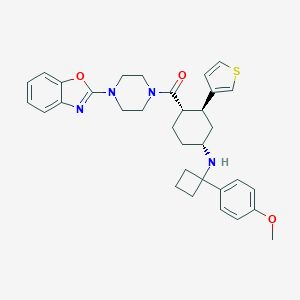 CHEMBL1940537 CHEMBL1940537 | C33H38N4O3S | 570.752 | 7 / 1 | 5.5 | No |
| 35402 | 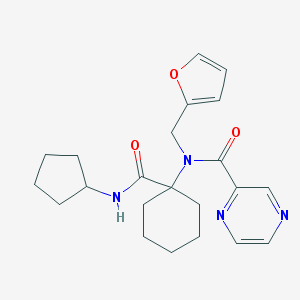 SMR000130166 SMR000130166 | C22H28N4O3 | 396.491 | 5 / 1 | 2.4 | Yes |
| 37139 | 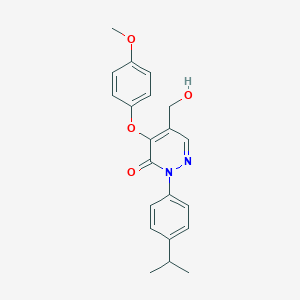 SR-02000000429 SR-02000000429 | C21H22N2O4 | 366.417 | 5 / 1 | 3.0 | Yes |
| 38806 | 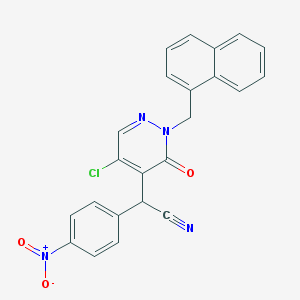 CHEMBL1988417 CHEMBL1988417 | C23H15ClN4O3 | 430.848 | 5 / 0 | 4.2 | Yes |
| 39434 | 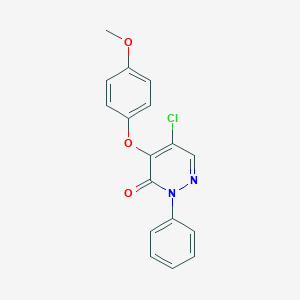 CHEMBL1441254 CHEMBL1441254 | C17H13ClN2O3 | 328.752 | 4 / 0 | 3.6 | Yes |
| 39657 | 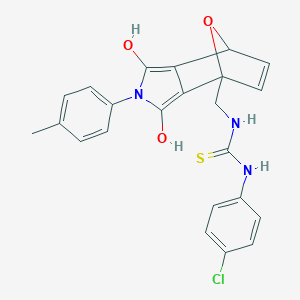 BDBM62096 BDBM62096 | C23H20ClN3O3S | 453.941 | 4 / 4 | 4.2 | Yes |
| 41360 | 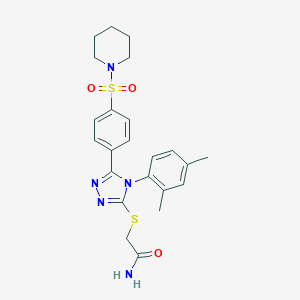 MLS000122826 MLS000122826 | C23H27N5O3S2 | 485.621 | 7 / 1 | 3.3 | Yes |
| 42115 | 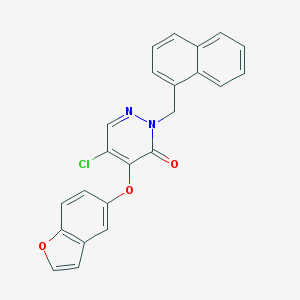 CHEMBL2314298 CHEMBL2314298 | C23H15ClN2O3 | 402.834 | 4 / 0 | 5.2 | No |
| 42362 | 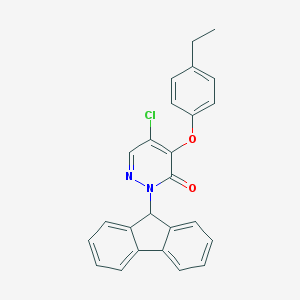 CHEMBL2314301 CHEMBL2314301 | C25H19ClN2O2 | 414.889 | 3 / 0 | 5.7 | No |
| 42715 | 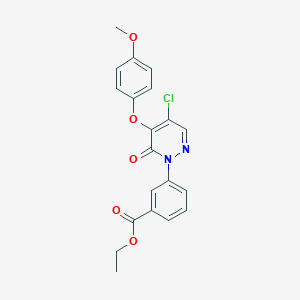 SR-02000000480 SR-02000000480 | C20H17ClN2O5 | 400.815 | 6 / 0 | 3.8 | Yes |
| 43746 | 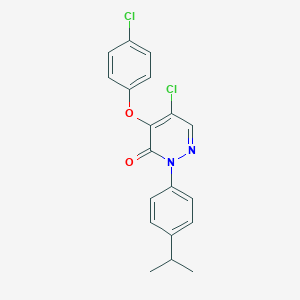 SR-02000000357 SR-02000000357 | C19H16Cl2N2O2 | 375.249 | 3 / 0 | 5.4 | No |
| 44721 | 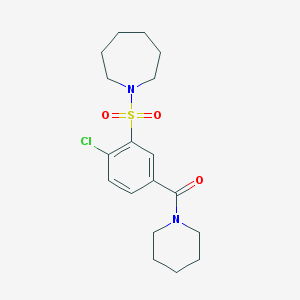 MLS001034823 MLS001034823 | C18H25ClN2O3S | 384.919 | 4 / 0 | 3.2 | Yes |
| 46324 | 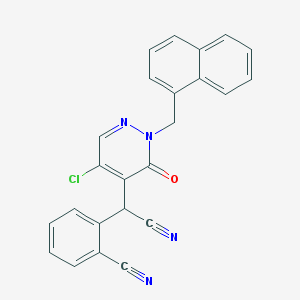 CHEMBL1982451 CHEMBL1982451 | C24H15ClN4O | 410.861 | 4 / 0 | 4.1 | Yes |
| 47114 | 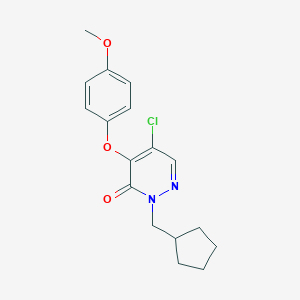 CHEMBL2314304 CHEMBL2314304 | C17H19ClN2O3 | 334.8 | 4 / 0 | 3.8 | Yes |
| 48620 | 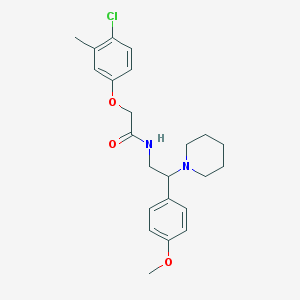 MLS001116287 MLS001116287 | C23H29ClN2O3 | 416.946 | 4 / 1 | 4.6 | Yes |
| 48780 | 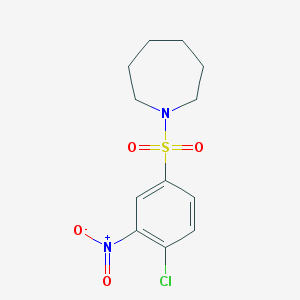 1-(4-Chloro-3-nitro-benzenesulfonyl)-azepane 1-(4-Chloro-3-nitro-benzenesulfonyl)-azepane | C12H15ClN2O4S | 318.772 | 5 / 0 | 2.7 | Yes |
| 49270 | 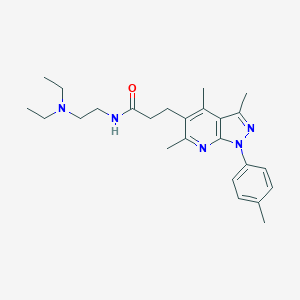 MLS001199098 MLS001199098 | C25H35N5O | 421.589 | 4 / 1 | 4.3 | Yes |
| 49507 | 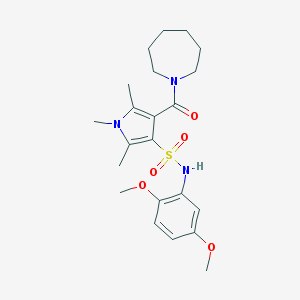 MLS001096203 MLS001096203 | C22H31N3O5S | 449.566 | 6 / 1 | 2.6 | Yes |
| 51354 | 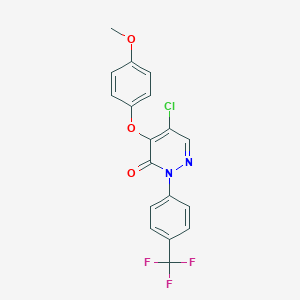 CHEMBL2207522 CHEMBL2207522 | C18H12ClF3N2O3 | 396.75 | 7 / 0 | 4.5 | Yes |
| 52658 | 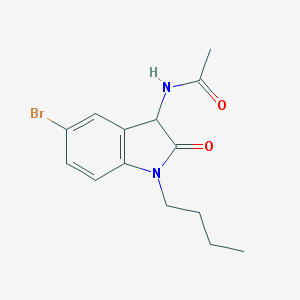 SMR000046229 SMR000046229 | C14H17BrN2O2 | 325.206 | 2 / 1 | 2.3 | Yes |
| 54304 | 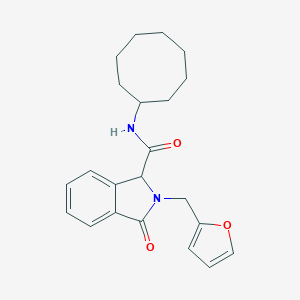 SMR000131465 SMR000131465 | C22H26N2O3 | 366.461 | 3 / 1 | 4.0 | Yes |
| 54762 | 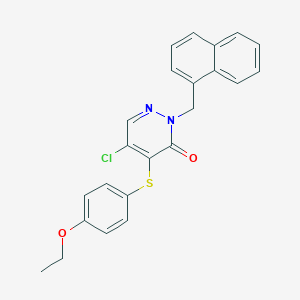 CHEMBL1997956 CHEMBL1997956 | C23H19ClN2O2S | 422.927 | 4 / 0 | 5.7 | No |
| 55063 | 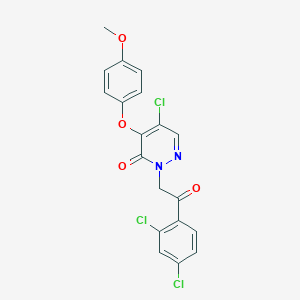 CHEMBL1992695 CHEMBL1992695 | C19H13Cl3N2O4 | 439.673 | 5 / 0 | 4.7 | Yes |
| 56641 | 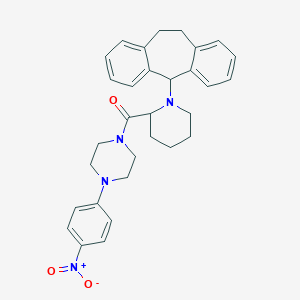 CHEMBL2068744 CHEMBL2068744 | C31H34N4O3 | 510.638 | 5 / 0 | 5.6 | No |
| 57580 | 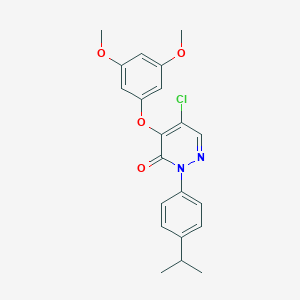 SR-02000000473 SR-02000000473 | C21H21ClN2O4 | 400.859 | 5 / 0 | 4.7 | Yes |
| 57674 | 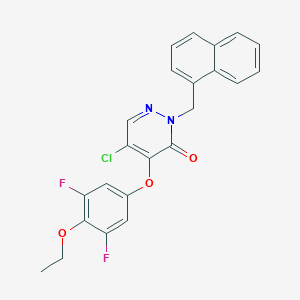 CHEMBL2002705 CHEMBL2002705 | C23H17ClF2N2O3 | 442.847 | 6 / 0 | 5.3 | No |
| 58164 | 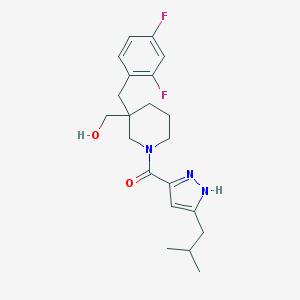 MLS001073551 MLS001073551 | C21H27F2N3O2 | 391.463 | 5 / 2 | 3.7 | Yes |
| 59213 | 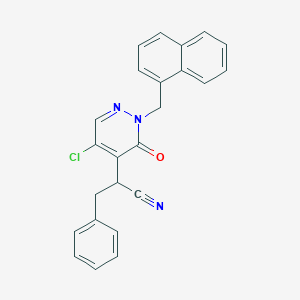 CHEMBL2001338 CHEMBL2001338 | C24H18ClN3O | 399.878 | 3 / 0 | 4.7 | Yes |
| 59958 | 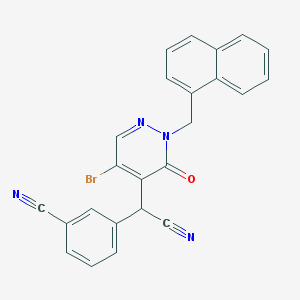 CHEMBL1992767 CHEMBL1992767 | C24H15BrN4O | 455.315 | 4 / 0 | 4.2 | Yes |
| 60433 | 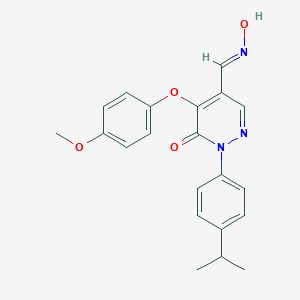 CHEMBL1729987 CHEMBL1729987 | C21H21N3O4 | 379.416 | 6 / 1 | 3.8 | Yes |
| 63555 | 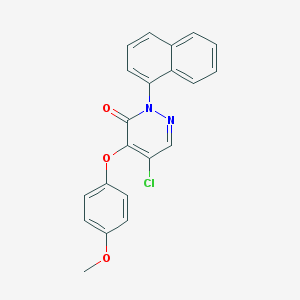 SR-02000000453 SR-02000000453 | C21H15ClN2O3 | 378.812 | 4 / 0 | 4.8 | Yes |
| 63825 | 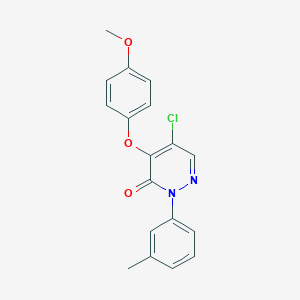 SR-02000000447 SR-02000000447 | C18H15ClN2O3 | 342.779 | 4 / 0 | 3.9 | Yes |
| 65279 | 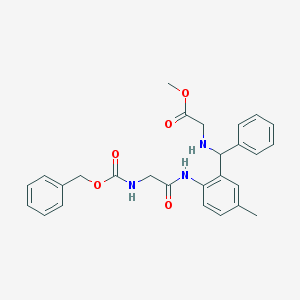 MLS000324743 MLS000324743 | C27H29N3O5 | 475.545 | 6 / 3 | 4.2 | Yes |
| 66697 | 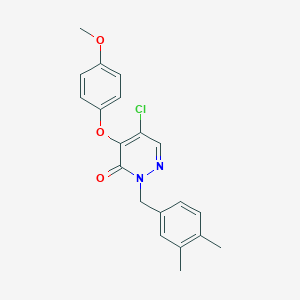 CHEMBL2314312 CHEMBL2314312 | C20H19ClN2O3 | 370.833 | 4 / 0 | 4.2 | Yes |
| 67087 | 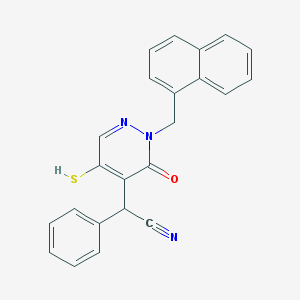 CHEMBL2003244 CHEMBL2003244 | C23H17N3OS | 383.469 | 4 / 1 | 3.9 | Yes |
| 67490 | 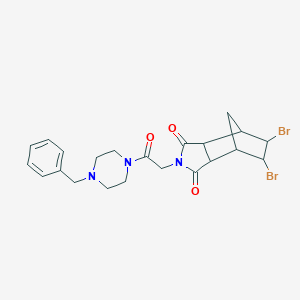 MLS000768518 MLS000768518 | C22H25Br2N3O3 | 539.268 | 4 / 0 | 2.1 | No |
| 559413 | 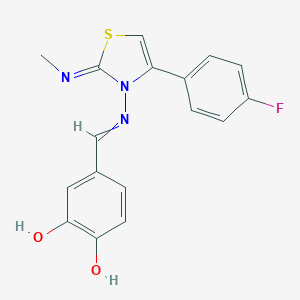 Oprea1_705406 Oprea1_705406 | C17H14FN3O2S | 343.376 | 6 / 2 | 3.3 | Yes |
| 71031 | 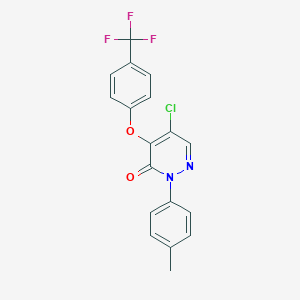 SR-02000000351 SR-02000000351 | C18H12ClF3N2O2 | 380.751 | 6 / 0 | 4.9 | Yes |
| 73942 | 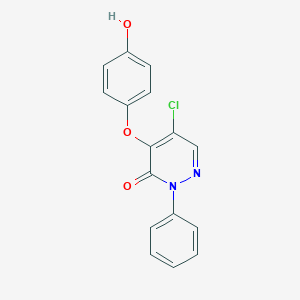 SR-02000000364 SR-02000000364 | C16H11ClN2O3 | 314.725 | 4 / 1 | 3.2 | Yes |
| 75202 | 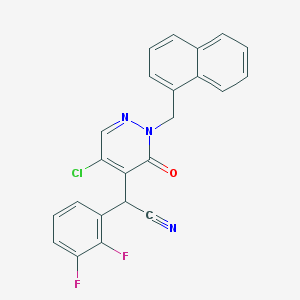 CHEMBL1964400 CHEMBL1964400 | C23H14ClF2N3O | 421.832 | 5 / 0 | 4.6 | Yes |
| 75432 | 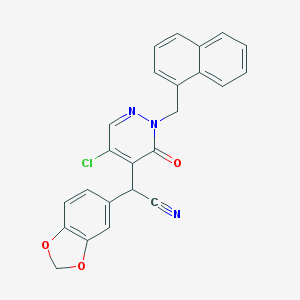 CHEMBL2004790 CHEMBL2004790 | C24H16ClN3O3 | 429.86 | 5 / 0 | 4.2 | Yes |
| 77100 | 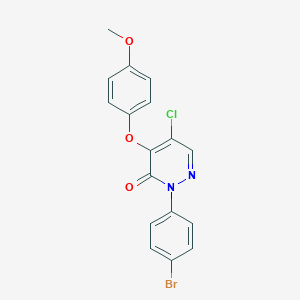 CHEMBL2207524 CHEMBL2207524 | C17H12BrClN2O3 | 407.648 | 4 / 0 | 4.3 | Yes |
| 77383 | 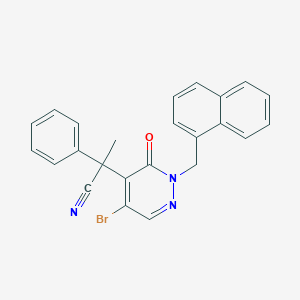 CHEMBL2004017 CHEMBL2004017 | C24H18BrN3O | 444.332 | 3 / 0 | 4.8 | Yes |
| 77489 | 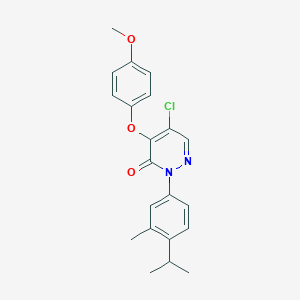 SR-02000000448 SR-02000000448 | C21H21ClN2O3 | 384.86 | 4 / 0 | 5.1 | No |
| 78435 | 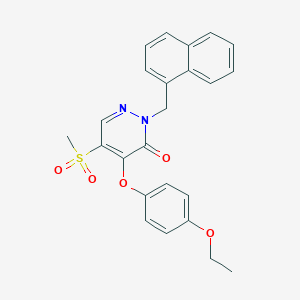 CHEMBL1997793 CHEMBL1997793 | C24H22N2O5S | 450.509 | 6 / 0 | 3.7 | Yes |
| 78947 | 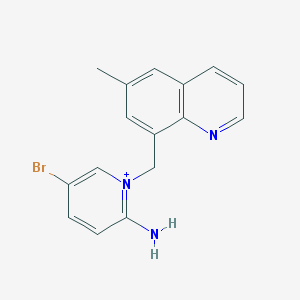 MLS001013428 MLS001013428 | C16H15BrN3+ | 329.221 | 2 / 1 | 3.6 | Yes |
| 79690 | 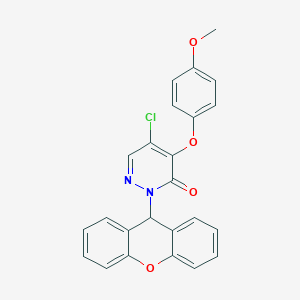 CHEMBL1973416 CHEMBL1973416 | C24H17ClN2O4 | 432.86 | 5 / 0 | 4.8 | Yes |
| 81227 | 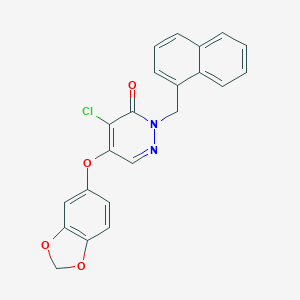 CHEMBL1985907 CHEMBL1985907 | C22H15ClN2O4 | 406.822 | 5 / 0 | 4.6 | Yes |
| 81487 | 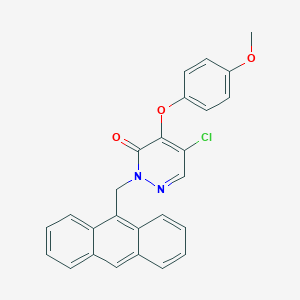 CHEMBL2314318 CHEMBL2314318 | C26H19ClN2O3 | 442.899 | 4 / 0 | 6.0 | No |
| 81882 | 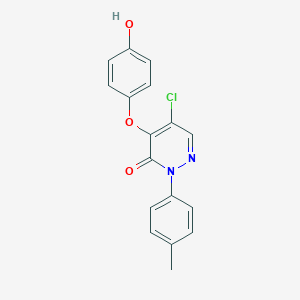 CHEMBL2207118 CHEMBL2207118 | C17H13ClN2O3 | 328.752 | 4 / 1 | 3.6 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218