

You can:
| Name | Histamine H4 receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Hrh4 |
| Synonym | GPCR105 H4 receptor H4R HH4R |
| Disease | N/A for non-human GPCRs |
| Length | 391 |
| Amino acid sequence | MSESNGTDVLPLTAQVPLAFLMSLLAFAITIGNAVVILAFVADRNLRHRSNYFFLNLAISDFFVGVISIPLYIPHTLFNWNFGSGICMFWLITDYLLCTASVYSIVLISYDRYQSVSNAVRYRAQHTGILKIVAQMVAVWILAFLVNGPMILASDSWKNSTNTEECEPGFVTEWYILAITAFLEFLLPVSLVVYFSVQIYWSLWKRGSLSRCPSHAGFIATSSRGTGHSRRTGLACRTSLPGLKEPAASLHSESPRGKSSLLVSLRTHMSGSIIAFKVGSFCRSESPVLHQREHVELLRGRKLARSLAVLLSAFAICWAPYCLFTIVLSTYRRGERPKSIWYSIAFWLQWFNSLINPFLYPLCHRRFQKAFWKILCVTKQPAPSQTQSVSS |
| UniProt | Q91ZY1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4468 |
| IUPHAR | 265 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 5512 | 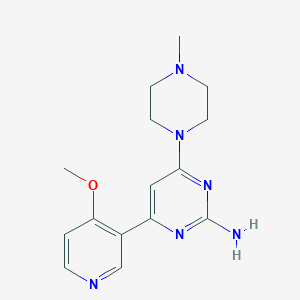 CHEMBL522019 CHEMBL522019 | C15H20N6O | 300.366 | 7 / 1 | 0.7 | Yes |
| 10510 | 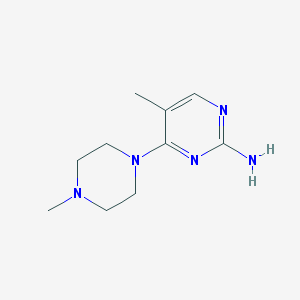 CHEMBL446896 CHEMBL446896 | C10H17N5 | 207.281 | 5 / 1 | 0.5 | Yes |
| 15064 | 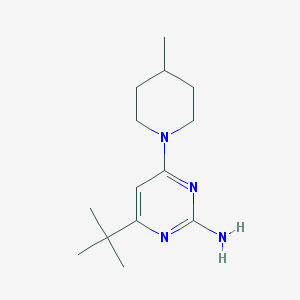 CHEMBL494687 CHEMBL494687 | C14H24N4 | 248.374 | 4 / 1 | 3.3 | Yes |
| 16340 | 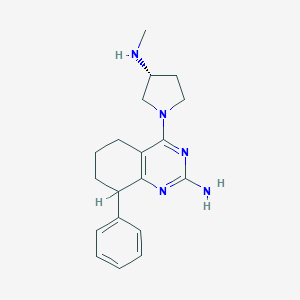 SCHEMBL605165 SCHEMBL605165 | C19H25N5 | 323.444 | 5 / 2 | 2.8 | Yes |
| 19307 | 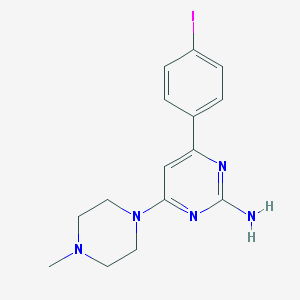 CHEMBL492288 CHEMBL492288 | C15H18IN5 | 395.248 | 5 / 1 | 2.4 | Yes |
| 24678 | 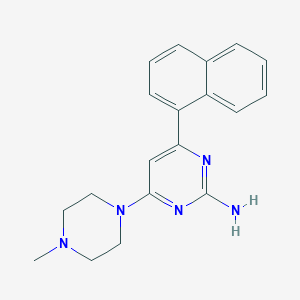 SCHEMBL607510 SCHEMBL607510 | C19H21N5 | 319.412 | 5 / 1 | 3.0 | Yes |
| 30140 | 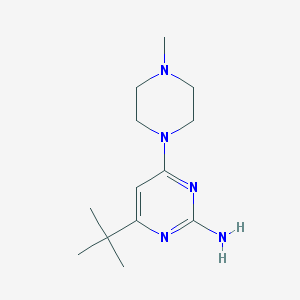 SCHEMBL604679 SCHEMBL604679 | C13H23N5 | 249.362 | 5 / 1 | 1.8 | Yes |
| 34367 | 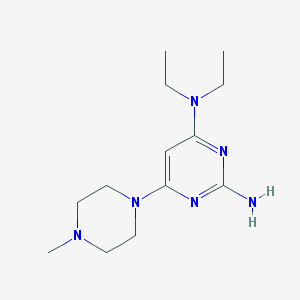 CHEMBL492662 CHEMBL492662 | C13H24N6 | 264.377 | 6 / 1 | 1.3 | Yes |
| 40267 | 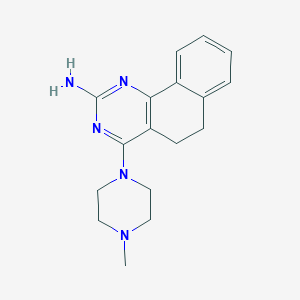 CHEMBL494093 CHEMBL494093 | C17H21N5 | 295.39 | 5 / 1 | 2.1 | Yes |
| 46822 | 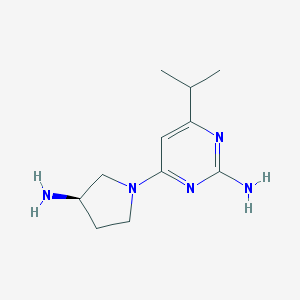 JNJ-39758979 JNJ-39758979 | C11H19N5 | 221.308 | 5 / 2 | 0.7 | Yes |
| 49757 | 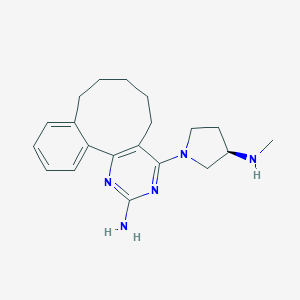 SCHEMBL606762 SCHEMBL606762 | C20H27N5 | 337.471 | 5 / 2 | 3.7 | Yes |
| 55478 | 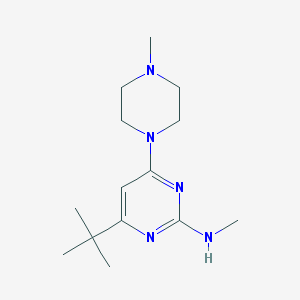 CHEMBL455016 CHEMBL455016 | C14H25N5 | 263.389 | 5 / 1 | 2.5 | Yes |
| 61554 | 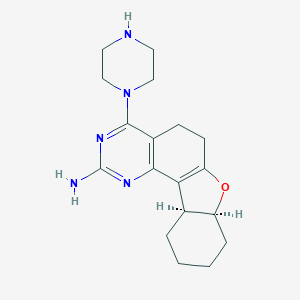 SCHEMBL604437 SCHEMBL604437 | C18H25N5O | 327.432 | 6 / 2 | 1.3 | Yes |
| 61658 | 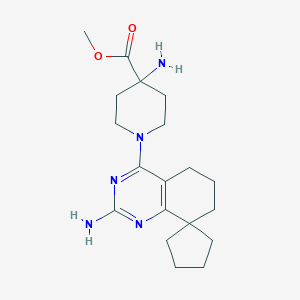 SCHEMBL603307 SCHEMBL603307 | C19H29N5O2 | 359.474 | 7 / 2 | 2.2 | Yes |
| 63094 | 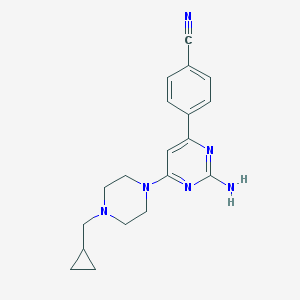 CHEMBL493885 CHEMBL493885 | C19H22N6 | 334.427 | 6 / 1 | 2.2 | Yes |
| 65969 | 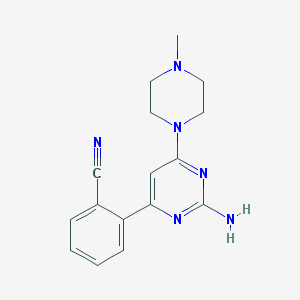 CHEMBL495083 CHEMBL495083 | C16H18N6 | 294.362 | 6 / 1 | 1.5 | Yes |
| 67182 | 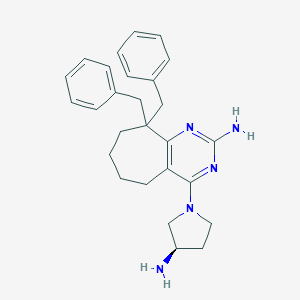 SCHEMBL606923 SCHEMBL606923 | C27H33N5 | 427.596 | 5 / 2 | 5.1 | No |
| 553659 | 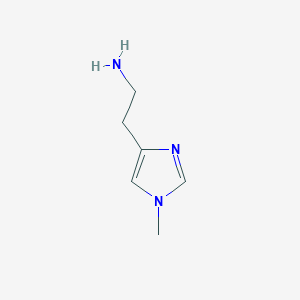 1-Methylhistamine 1-Methylhistamine | C6H11N3 | 125.175 | 2 / 1 | -0.6 | Yes |
| 81014 | 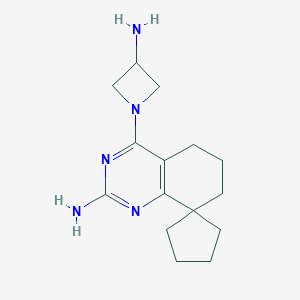 SCHEMBL604850 SCHEMBL604850 | C15H23N5 | 273.384 | 5 / 2 | 1.8 | Yes |
| 82707 | 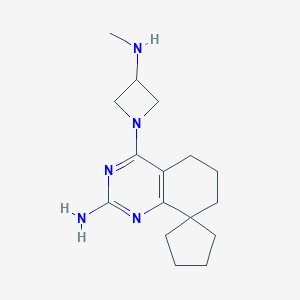 CHEMBL1079516 CHEMBL1079516 | C16H25N5 | 287.411 | 5 / 2 | 2.3 | Yes |
| 88287 | 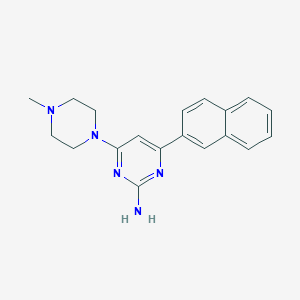 CHEMBL522841 CHEMBL522841 | C19H21N5 | 319.412 | 5 / 1 | 3.0 | Yes |
| 95580 | 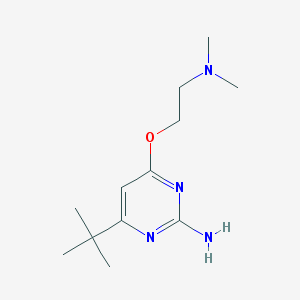 CHEMBL495341 CHEMBL495341 | C12H22N4O | 238.335 | 5 / 1 | 1.9 | Yes |
| 96439 | 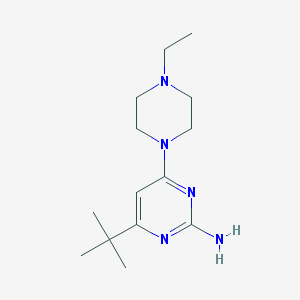 CHEMBL506187 CHEMBL506187 | C14H25N5 | 263.389 | 5 / 1 | 2.2 | Yes |
| 99323 | 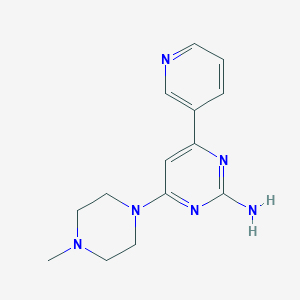 CHEMBL492227 CHEMBL492227 | C14H18N6 | 270.34 | 6 / 1 | 0.7 | Yes |
| 101431 | 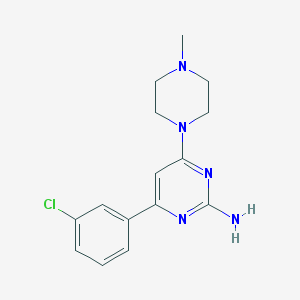 CHEMBL492287 CHEMBL492287 | C15H18ClN5 | 303.794 | 5 / 1 | 2.4 | Yes |
| 104032 | 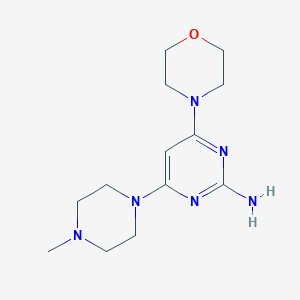 CHEMBL444484 CHEMBL444484 | C13H22N6O | 278.36 | 7 / 1 | 0.2 | Yes |
| 107817 | 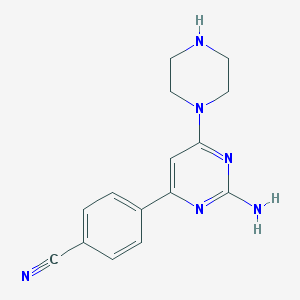 JMC516547 Compound 5 JMC516547 Compound 5 | C15H16N6 | 280.335 | 6 / 2 | 1.0 | Yes |
| 107860 | 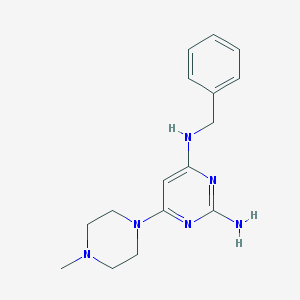 CHEMBL492677 CHEMBL492677 | C16H22N6 | 298.394 | 6 / 2 | 1.9 | Yes |
| 108441 | 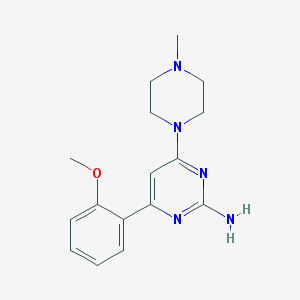 CHEMBL493719 CHEMBL493719 | C16H21N5O | 299.378 | 6 / 1 | 1.8 | Yes |
| 110857 | 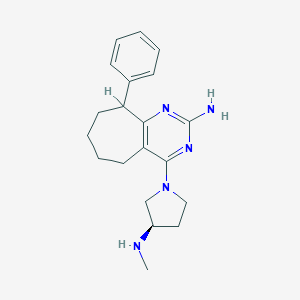 SCHEMBL605799 SCHEMBL605799 | C20H27N5 | 337.471 | 5 / 2 | 3.3 | Yes |
| 112622 | 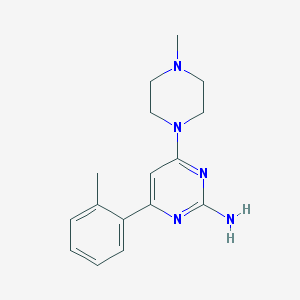 CHEMBL494723 CHEMBL494723 | C16H21N5 | 283.379 | 5 / 1 | 2.1 | Yes |
| 115041 | 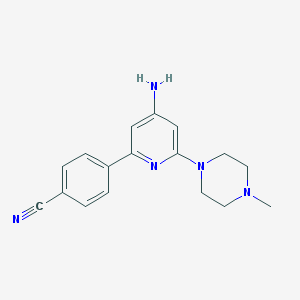 CHEMBL493462 CHEMBL493462 | C17H19N5 | 293.374 | 5 / 1 | 1.8 | Yes |
| 115286 | 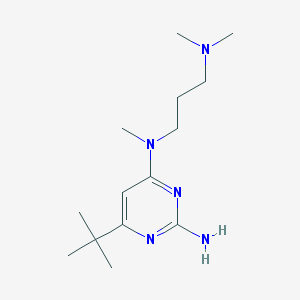 CHEMBL493100 CHEMBL493100 | C14H27N5 | 265.405 | 5 / 1 | 2.4 | Yes |
| 120386 | 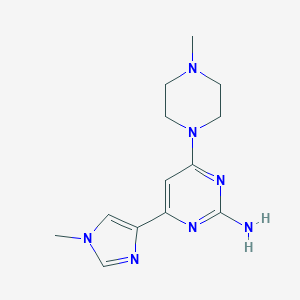 CHEMBL494576 CHEMBL494576 | C13H19N7 | 273.344 | 6 / 1 | -0.1 | Yes |
| 121546 | 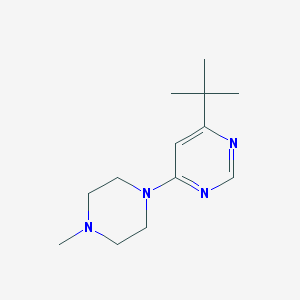 CHEMBL495342 CHEMBL495342 | C13H22N4 | 234.347 | 4 / 0 | 2.2 | Yes |
| 123064 | 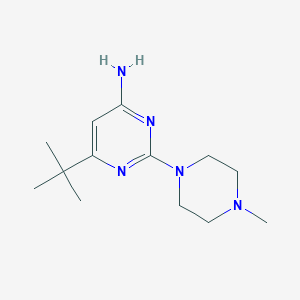 CHEMBL493261 CHEMBL493261 | C13H23N5 | 249.362 | 5 / 1 | 1.8 | Yes |
| 123587 | 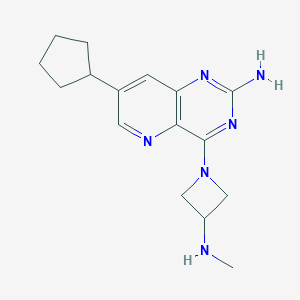 CHEMBL2376803 CHEMBL2376803 | C16H22N6 | 298.394 | 6 / 2 | 2.1 | Yes |
| 124516 | 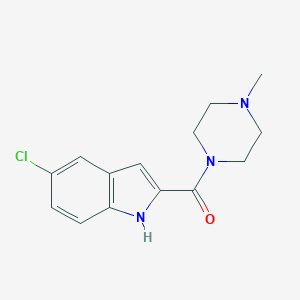 JNJ-7777120 JNJ-7777120 | C14H16ClN3O | 277.752 | 2 / 1 | 2.3 | Yes |
| 125399 | 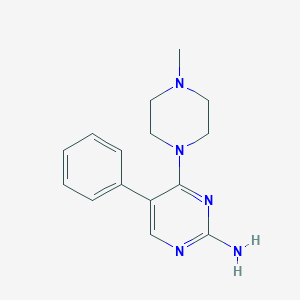 CHEMBL451108 CHEMBL451108 | C15H19N5 | 269.352 | 5 / 1 | 1.7 | Yes |
| 126248 | 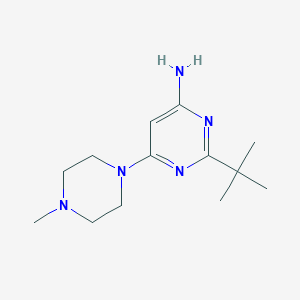 CHEMBL489663 CHEMBL489663 | C13H23N5 | 249.362 | 5 / 1 | 1.8 | Yes |
| 127226 | 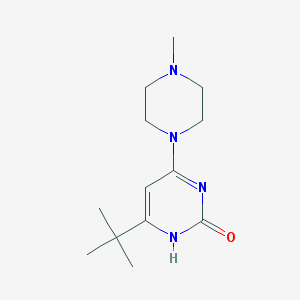 CHEMBL447151 CHEMBL447151 | C13H22N4O | 250.346 | 2 / 1 | 0.8 | Yes |
| 129128 | 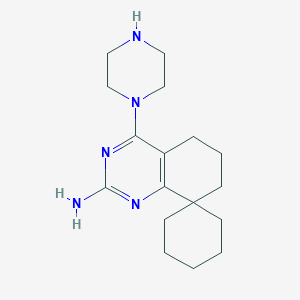 CHEMBL1081683 CHEMBL1081683 | C17H27N5 | 301.438 | 5 / 2 | 2.8 | Yes |
| 134789 | 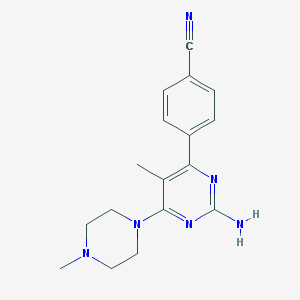 CHEMBL492900 CHEMBL492900 | C17H20N6 | 308.389 | 6 / 1 | 1.9 | Yes |
| 136157 | 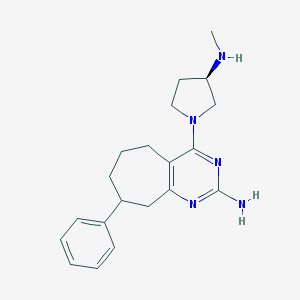 CHEMBL1089040 CHEMBL1089040 | C20H27N5 | 337.471 | 5 / 2 | 3.2 | Yes |
| 136932 | 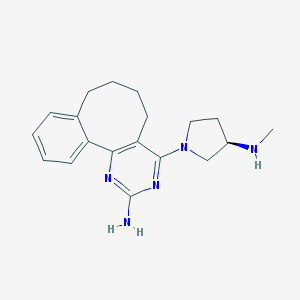 SCHEMBL606075 SCHEMBL606075 | C19H25N5 | 323.444 | 5 / 2 | 3.1 | Yes |
| 137225 | 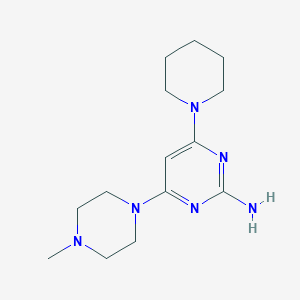 SCHEMBL605178 SCHEMBL605178 | C14H24N6 | 276.388 | 6 / 1 | 1.4 | Yes |
| 137572 | 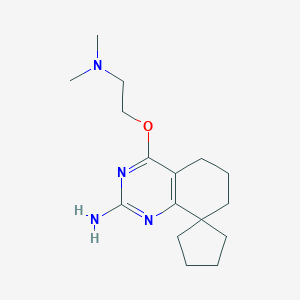 SCHEMBL604978 SCHEMBL604978 | C16H26N4O | 290.411 | 5 / 1 | 2.8 | Yes |
| 139175 | 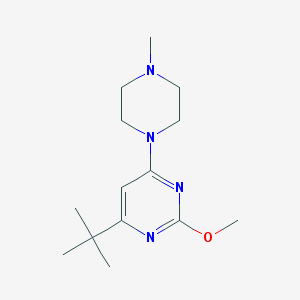 CHEMBL492489 CHEMBL492489 | C14H24N4O | 264.373 | 5 / 0 | 2.5 | Yes |
| 140556 | 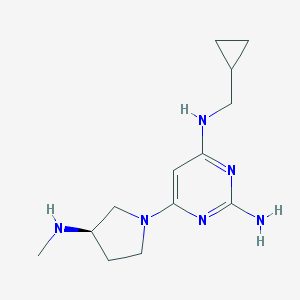 PF-3893787 PF-3893787 | C13H22N6 | 262.361 | 6 / 3 | 1.1 | Yes |
| 140713 | 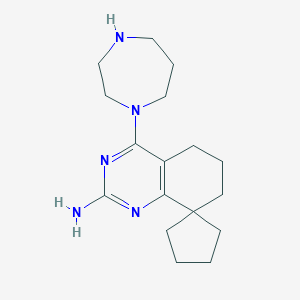 SCHEMBL604462 SCHEMBL604462 | C17H27N5 | 301.438 | 5 / 2 | 2.6 | Yes |
| 141572 | 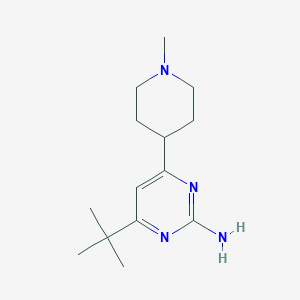 CHEMBL494585 CHEMBL494585 | C14H24N4 | 248.374 | 4 / 1 | 2.2 | Yes |
| 150353 | 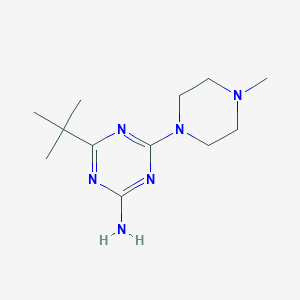 CHEMBL494679 CHEMBL494679 | C12H22N6 | 250.35 | 6 / 1 | 1.6 | Yes |
| 153370 | 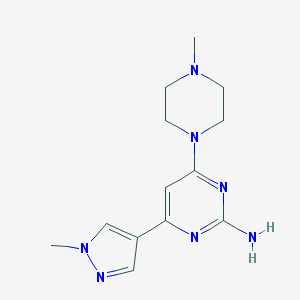 CHEMBL493229 CHEMBL493229 | C13H19N7 | 273.344 | 6 / 1 | 0.1 | Yes |
| 154963 | 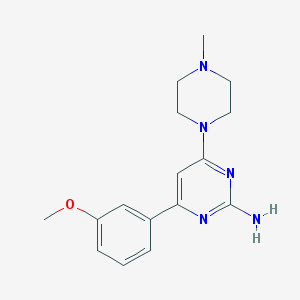 CHEMBL494724 CHEMBL494724 | C16H21N5O | 299.378 | 6 / 1 | 1.8 | Yes |
| 155372 | 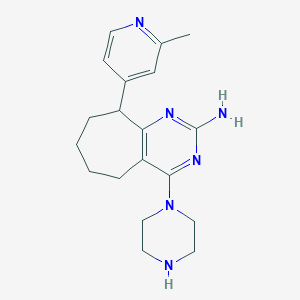 SCHEMBL604979 SCHEMBL604979 | C19H26N6 | 338.459 | 6 / 2 | 2.2 | Yes |
| 158443 | 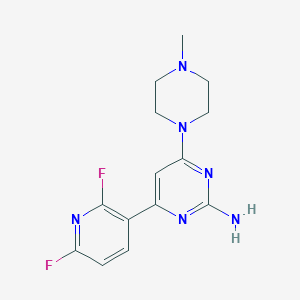 CHEMBL492242 CHEMBL492242 | C14H16F2N6 | 306.321 | 8 / 1 | 1.6 | Yes |
| 161525 | 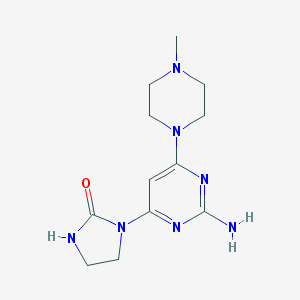 CHEMBL493663 CHEMBL493663 | C12H19N7O | 277.332 | 6 / 2 | -0.6 | Yes |
| 163262 | 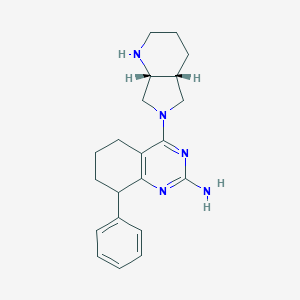 SCHEMBL602901 SCHEMBL602901 | C21H27N5 | 349.482 | 5 / 2 | 3.1 | Yes |
| 166281 | 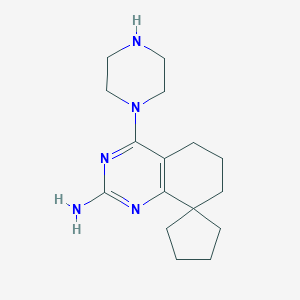 SCHEMBL606187 SCHEMBL606187 | C16H25N5 | 287.411 | 5 / 2 | 2.3 | Yes |
| 168057 | 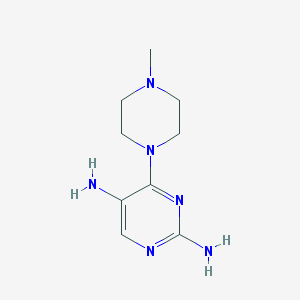 CHEMBL493478 CHEMBL493478 | C9H16N6 | 208.269 | 6 / 2 | -0.6 | Yes |
| 169716 | 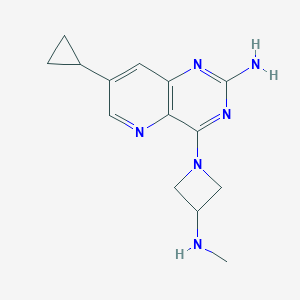 CHEMBL2376804 CHEMBL2376804 | C14H18N6 | 270.34 | 6 / 2 | 1.0 | Yes |
| 172196 | 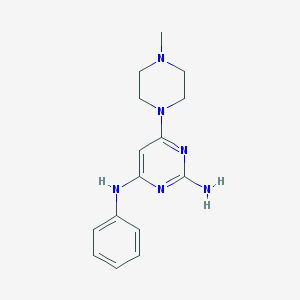 CHEMBL502104 CHEMBL502104 | C15H20N6 | 284.367 | 6 / 2 | 2.0 | Yes |
| 173140 | 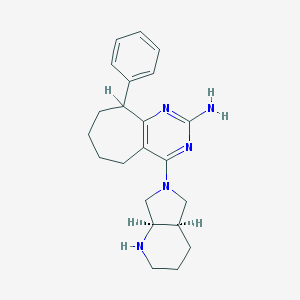 SCHEMBL4054703 SCHEMBL4054703 | C22H29N5 | 363.509 | 5 / 2 | 3.7 | Yes |
| 173141 | 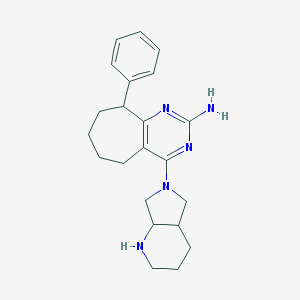 CHEMBL1077276 CHEMBL1077276 | C22H29N5 | 363.509 | 5 / 2 | 3.7 | Yes |
| 177298 | 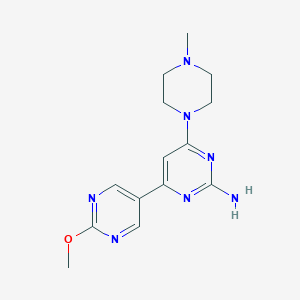 CHEMBL523835 CHEMBL523835 | C14H19N7O | 301.354 | 8 / 1 | 0.4 | Yes |
| 177754 | 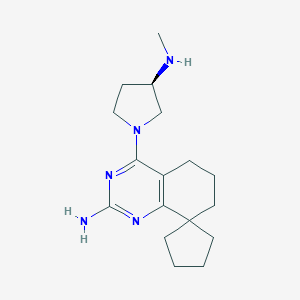 CHEMBL1089390 CHEMBL1089390 | C17H27N5 | 301.438 | 5 / 2 | 2.7 | Yes |
| 178949 | 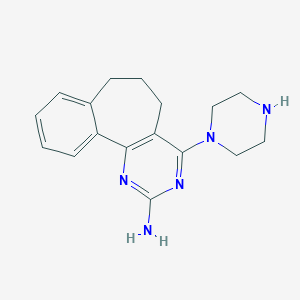 UNII-8U5620H545 UNII-8U5620H545 | C17H21N5 | 295.39 | 5 / 2 | 2.2 | Yes |
| 181841 | 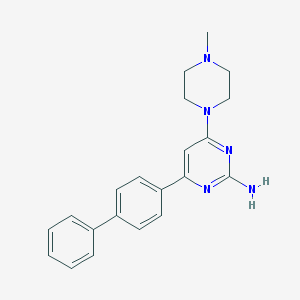 CHEMBL523361 CHEMBL523361 | C21H23N5 | 345.45 | 5 / 1 | 3.4 | Yes |
| 182589 | 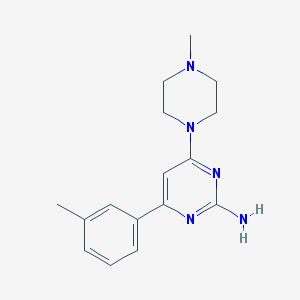 CHEMBL492274 CHEMBL492274 | C16H21N5 | 283.379 | 5 / 1 | 2.1 | Yes |
| 183131 | 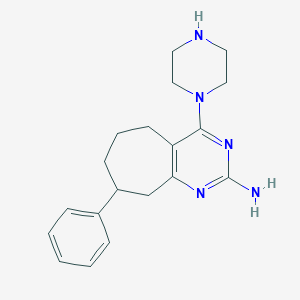 SCHEMBL607364 SCHEMBL607364 | C19H25N5 | 323.444 | 5 / 2 | 2.8 | Yes |
| 192749 | 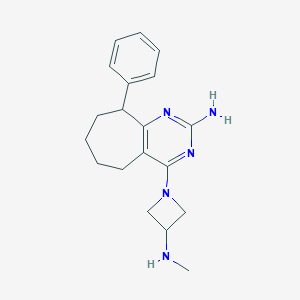 SCHEMBL606041 SCHEMBL606041 | C19H25N5 | 323.444 | 5 / 2 | 2.9 | Yes |
| 194047 | 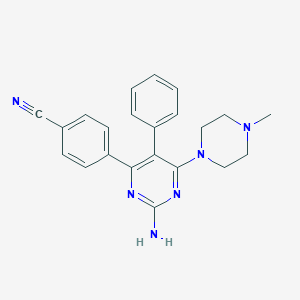 CHEMBL493104 CHEMBL493104 | C22H22N6 | 370.46 | 6 / 1 | 3.1 | Yes |
| 196193 | 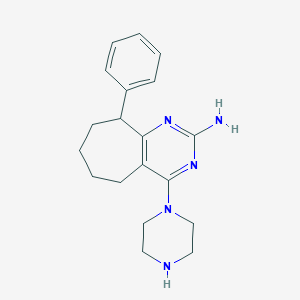 SCHEMBL606676 SCHEMBL606676 | C19H25N5 | 323.444 | 5 / 2 | 2.9 | Yes |
| 197072 | 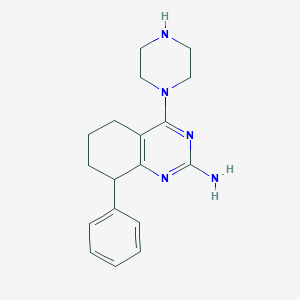 SCHEMBL603530 SCHEMBL603530 | C18H23N5 | 309.417 | 5 / 2 | 2.3 | Yes |
| 202306 | 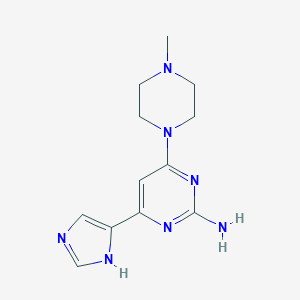 CHEMBL494667 CHEMBL494667 | C12H17N7 | 259.317 | 6 / 2 | 0.0 | Yes |
| 563985 | 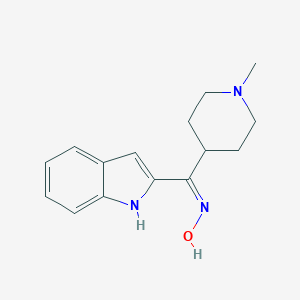 CHEMBL1632411 CHEMBL1632411 | C15H19N3O | 257.337 | 3 / 2 | 2.8 | Yes |
| 224077 | 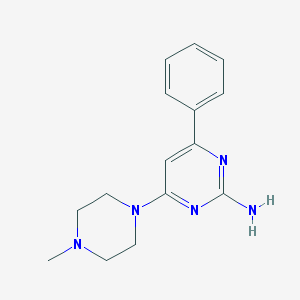 VUF10460 VUF10460 | C15H19N5 | 269.352 | 5 / 1 | 1.8 | Yes |
| 224250 | 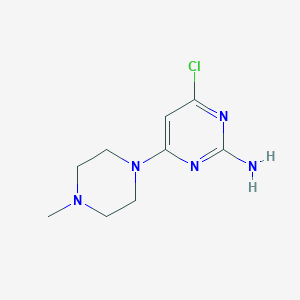 4-chloro-6-(4-methylpiperazin-1-yl)pyrimidin-2-amine 4-chloro-6-(4-methylpiperazin-1-yl)pyrimidin-2-amine | C9H14ClN5 | 227.696 | 5 / 1 | 1.1 | Yes |
| 225856 | 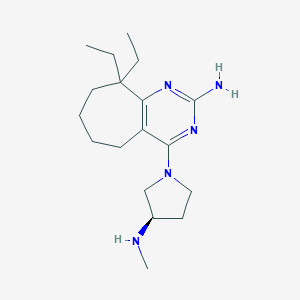 CHEMBL1092148 CHEMBL1092148 | C18H31N5 | 317.481 | 5 / 2 | 3.4 | Yes |
| 226482 | 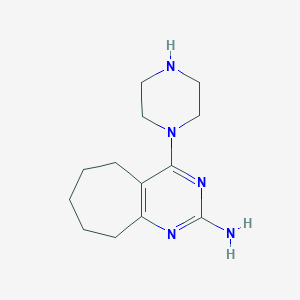 SCHEMBL605565 SCHEMBL605565 | C13H21N5 | 247.346 | 5 / 2 | 1.4 | Yes |
| 228126 | 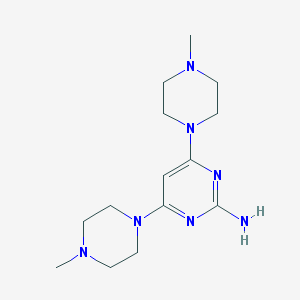 CHEMBL492869 CHEMBL492869 | C14H25N7 | 291.403 | 7 / 1 | 0.4 | Yes |
| 232087 | 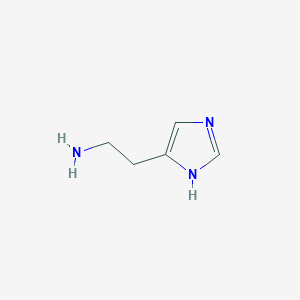 histamine histamine | C5H9N3 | 111.148 | 2 / 2 | -0.7 | Yes |
| 232513 | 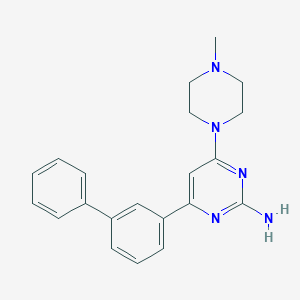 CHEMBL494070 CHEMBL494070 | C21H23N5 | 345.45 | 5 / 1 | 3.4 | Yes |
| 235789 | 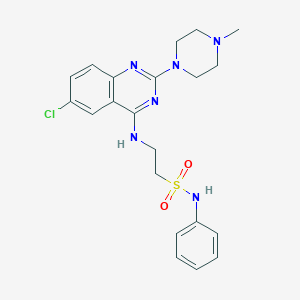 CHEMBL592379 CHEMBL592379 | C21H25ClN6O2S | 460.981 | 8 / 2 | 4.0 | Yes |
| 240392 | 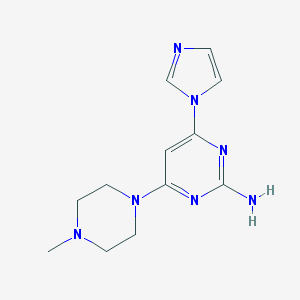 CHEMBL448085 CHEMBL448085 | C12H17N7 | 259.317 | 6 / 1 | 0.2 | Yes |
| 241255 | 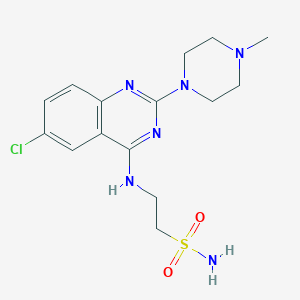 CHEMBL591969 CHEMBL591969 | C15H21ClN6O2S | 384.883 | 8 / 2 | 2.0 | Yes |
| 242784 | 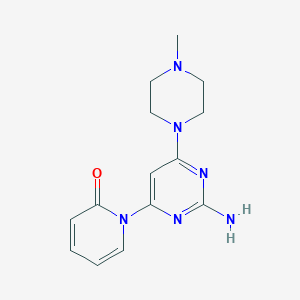 CHEMBL493664 CHEMBL493664 | C14H18N6O | 286.339 | 6 / 1 | 0.5 | Yes |
| 244358 | 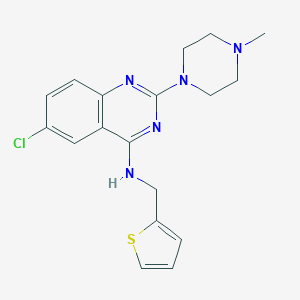 CHEMBL452847 CHEMBL452847 | C18H20ClN5S | 373.903 | 6 / 1 | 4.0 | Yes |
| 246187 | 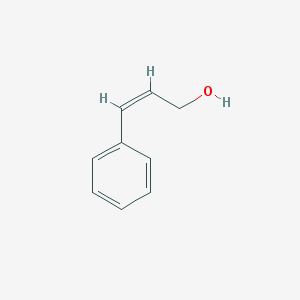 Z-cinnamyl alcohol Z-cinnamyl alcohol | C9H10O | 134.178 | 1 / 1 | 1.9 | Yes |
| 246188 | 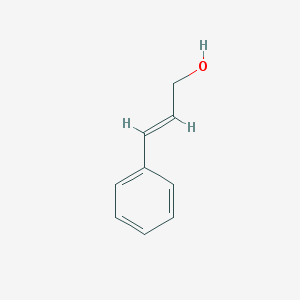 cinnamyl alcohol cinnamyl alcohol | C9H10O | 134.178 | 1 / 1 | 1.9 | Yes |
| 248322 | 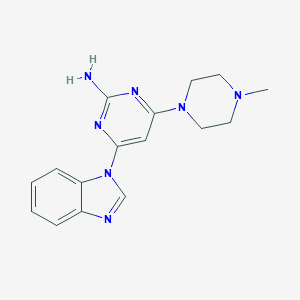 CHEMBL495280 CHEMBL495280 | C16H19N7 | 309.377 | 6 / 1 | 1.6 | Yes |
| 248399 | 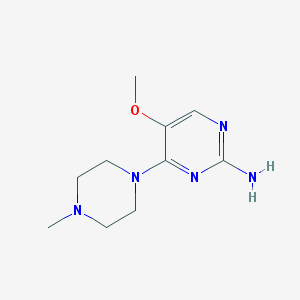 CHEMBL493477 CHEMBL493477 | C10H17N5O | 223.28 | 6 / 1 | 0.1 | Yes |
| 255033 | 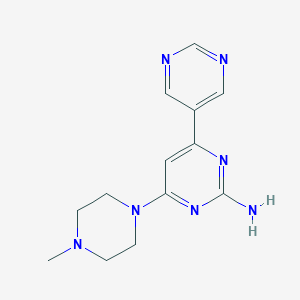 CHEMBL505346 CHEMBL505346 | C13H17N7 | 271.328 | 7 / 1 | 0.1 | Yes |
| 554554 | 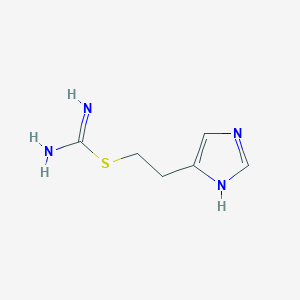 imetit imetit | C6H10N4S | 170.234 | 3 / 3 | 0.2 | Yes |
| 266829 | 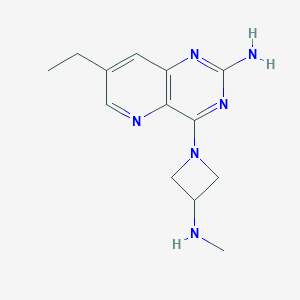 CHEMBL2376800 CHEMBL2376800 | C13H18N6 | 258.329 | 6 / 2 | 0.9 | Yes |
| 268237 | 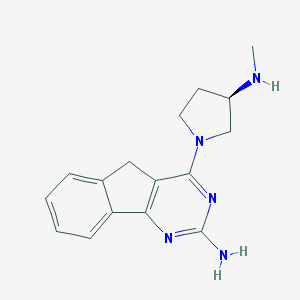 CHEMBL446269 CHEMBL446269 | C16H19N5 | 281.363 | 5 / 2 | 1.7 | Yes |
| 273703 | 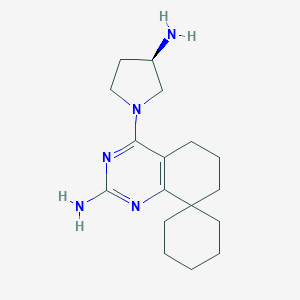 CHEMBL1078018 CHEMBL1078018 | C17H27N5 | 301.438 | 5 / 2 | 2.7 | Yes |
| 274790 | 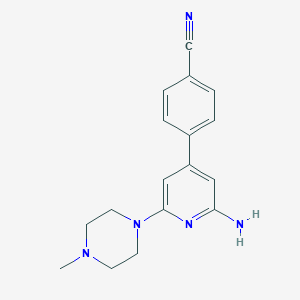 CHEMBL493262 CHEMBL493262 | C17H19N5 | 293.374 | 5 / 1 | 2.1 | Yes |
| 276032 | 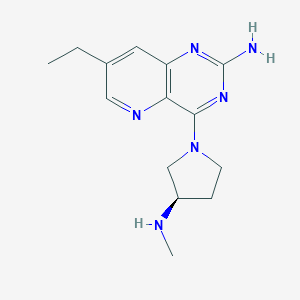 CHEMBL2376801 CHEMBL2376801 | C14H20N6 | 272.356 | 6 / 2 | 1.2 | Yes |
| 276961 | 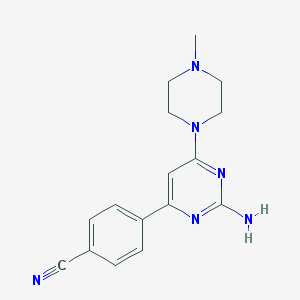 A-846714 A-846714 | C16H18N6 | 294.362 | 6 / 1 | 1.5 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218